Chapter 6 Multigroup Confirmatory Factor Analysis
6.1 Syntax - R
6.1.1 Analysis with data from single groups
NELS.MALE.corr <- '
1
0.331 1
0.277 0.416 1
0.482 0.37 0.31 1
0.203 0.162 0.141 0.227 1
0.057 0.161 0.096 0.123 0.242 1
0.207 0.144 0.137 0.236 0.302 0.196 1
0.211 0.188 0.172 0.26 0.293 0.235 0.401 1'
NELS.MALE.SDs <- c(0.586, 0.643, 0.620, 0.653, 0.804, 0.748, 0.763, 0.766)
NELS.FEMALE.corr <- '
1
0.418 1
0.34 0.439 1
0.581 0.431 0.364 1
0.223 0.209 0.169 0.282 1
0.075 0.158 0.121 0.155 0.276 1
0.233 0.197 0.192 0.281 0.338 0.231 1
0.269 0.249 0.215 0.333 0.358 0.251 0.465 1'
NELS.FEMALE.SDs <- c(0.616, 0.650, 0.642, 0.693, 0.805, 0.703, 0.754, 0.796)
NELS.MALE.cov <- getCov(NELS.MALE.corr, sds = NELS.MALE.SDs, names = c(paste0('byconf', 1:4, sep=""), paste0('byloc', 1:4, sep="")))
NELS.FEMALE.cov <- getCov(NELS.FEMALE.corr, sds = NELS.FEMALE.SDs, names = c(paste0('byconf', 1:4, sep=""), paste0('byloc', 1:4, sep="")))
NELS.cfa.model <-
paste0('f1 =~ ', paste0('byconf', 1:4, collapse=' + '), ' \n',
' f2 =~ ', paste0('byloc', 1:4, collapse=' + '), ' \n',
' byconf2 ~~ byconf3')
NELS.MALE.fit <- cfa(NELS.cfa.model, sample.cov = NELS.MALE.cov, sample.nobs = 5312)
NELS.FEMALE.fit <- cfa(NELS.cfa.model, sample.cov = NELS.FEMALE.cov, sample.nobs = 6008)
summary(NELS.MALE.fit, fit.measures = TRUE, standardized = TRUE, rsquare = TRUE) ## lavaan 0.6-8 ended normally after 33 iterations
##
## Estimator ML
## Optimization method NLMINB
## Number of model parameters 18
##
## Number of observations 5312
##
## Model Test User Model:
##
## Test statistic 146.970
## Degrees of freedom 18
## P-value (Chi-square) 0.000
##
## Model Test Baseline Model:
##
## Test statistic 6651.699
## Degrees of freedom 28
## P-value 0.000
##
## User Model versus Baseline Model:
##
## Comparative Fit Index (CFI) 0.981
## Tucker-Lewis Index (TLI) 0.970
##
## Loglikelihood and Information Criteria:
##
## Loglikelihood user model (H0) -41500.792
## Loglikelihood unrestricted model (H1) -41427.307
##
## Akaike (AIC) 83037.584
## Bayesian (BIC) 83155.983
## Sample-size adjusted Bayesian (BIC) 83098.785
##
## Root Mean Square Error of Approximation:
##
## RMSEA 0.037
## 90 Percent confidence interval - lower 0.031
## 90 Percent confidence interval - upper 0.042
## P-value RMSEA <= 0.05 1.000
##
## Standardized Root Mean Square Residual:
##
## SRMR 0.022
##
## Parameter Estimates:
##
## Standard errors Standard
## Information Expected
## Information saturated (h1) model Structured
##
## Latent Variables:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## f1 =~
## byconf1 1.000 0.379 0.647
## byconf2 0.863 0.031 27.486 0.000 0.327 0.509
## byconf3 0.701 0.029 23.820 0.000 0.266 0.428
## byconf4 1.272 0.040 31.424 0.000 0.482 0.738
## f2 =~
## byloc1 1.000 0.412 0.512
## byloc2 0.667 0.036 18.716 0.000 0.274 0.367
## byloc3 1.115 0.045 24.712 0.000 0.459 0.602
## byloc4 1.179 0.047 24.971 0.000 0.485 0.634
##
## Covariances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .byconf2 ~~
## .byconf3 0.079 0.005 15.304 0.000 0.079 0.255
## f1 ~~
## f2 0.085 0.004 18.978 0.000 0.546 0.546
##
## Variances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .byconf1 0.200 0.006 35.342 0.000 0.200 0.582
## .byconf2 0.306 0.007 43.856 0.000 0.306 0.741
## .byconf3 0.314 0.007 46.432 0.000 0.314 0.817
## .byconf4 0.194 0.007 25.998 0.000 0.194 0.455
## .byloc1 0.477 0.011 42.099 0.000 0.477 0.738
## .byloc2 0.484 0.010 47.493 0.000 0.484 0.865
## .byloc3 0.371 0.010 36.188 0.000 0.371 0.638
## .byloc4 0.351 0.010 33.480 0.000 0.351 0.599
## f1 0.144 0.007 21.119 0.000 1.000 1.000
## f2 0.169 0.011 15.846 0.000 1.000 1.000
##
## R-Square:
## Estimate
## byconf1 0.418
## byconf2 0.259
## byconf3 0.183
## byconf4 0.545
## byloc1 0.262
## byloc2 0.135
## byloc3 0.362
## byloc4 0.401## lavaan 0.6-8 ended normally after 31 iterations
##
## Estimator ML
## Optimization method NLMINB
## Number of model parameters 18
##
## Number of observations 6008
##
## Model Test User Model:
##
## Test statistic 198.307
## Degrees of freedom 18
## P-value (Chi-square) 0.000
##
## Model Test Baseline Model:
##
## Test statistic 10198.456
## Degrees of freedom 28
## P-value 0.000
##
## User Model versus Baseline Model:
##
## Comparative Fit Index (CFI) 0.982
## Tucker-Lewis Index (TLI) 0.972
##
## Loglikelihood and Information Criteria:
##
## Loglikelihood user model (H0) -46343.263
## Loglikelihood unrestricted model (H1) -46244.110
##
## Akaike (AIC) 92722.527
## Bayesian (BIC) 92843.142
## Sample-size adjusted Bayesian (BIC) 92785.943
##
## Root Mean Square Error of Approximation:
##
## RMSEA 0.041
## 90 Percent confidence interval - lower 0.036
## 90 Percent confidence interval - upper 0.046
## P-value RMSEA <= 0.05 0.998
##
## Standardized Root Mean Square Residual:
##
## SRMR 0.024
##
## Parameter Estimates:
##
## Standard errors Standard
## Information Expected
## Information saturated (h1) model Structured
##
## Latent Variables:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## f1 =~
## byconf1 1.000 0.445 0.722
## byconf2 0.820 0.022 36.510 0.000 0.365 0.561
## byconf3 0.677 0.022 30.855 0.000 0.301 0.469
## byconf4 1.240 0.028 43.702 0.000 0.552 0.796
## f2 =~
## byloc1 1.000 0.441 0.548
## byloc2 0.604 0.027 22.120 0.000 0.267 0.379
## byloc3 1.091 0.035 30.827 0.000 0.482 0.639
## byloc4 1.261 0.040 31.500 0.000 0.556 0.699
##
## Covariances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .byconf2 ~~
## .byconf3 0.073 0.005 15.686 0.000 0.073 0.240
## f1 ~~
## f2 0.112 0.005 23.313 0.000 0.571 0.571
##
## Variances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .byconf1 0.181 0.005 36.091 0.000 0.181 0.478
## .byconf2 0.289 0.006 47.312 0.000 0.289 0.685
## .byconf3 0.321 0.006 50.052 0.000 0.321 0.780
## .byconf4 0.176 0.007 26.934 0.000 0.176 0.366
## .byloc1 0.453 0.010 45.301 0.000 0.453 0.700
## .byloc2 0.423 0.008 51.184 0.000 0.423 0.856
## .byloc3 0.336 0.009 39.152 0.000 0.336 0.592
## .byloc4 0.324 0.010 33.420 0.000 0.324 0.512
## f1 0.198 0.007 27.642 0.000 1.000 1.000
## f2 0.195 0.010 19.176 0.000 1.000 1.000
##
## R-Square:
## Estimate
## byconf1 0.522
## byconf2 0.315
## byconf3 0.220
## byconf4 0.634
## byloc1 0.300
## byloc2 0.144
## byloc3 0.408
## byloc4 0.4886.1.2 Analysis with data from both groups
For multiple group analysis, “sample.cov” should be a list with a variance-covariance matrix of each group, and “sample.nobs” should be a list or a vector with the number of observations for each group. By default, the same model is fitted in all groups without any equality constraints on the model parameters
6.1.2.1 both groups, no constraints
NELS.twogroup.fit <- cfa(NELS.cfa.model, sample.cov = list(NELS.MALE.cov, NELS.FEMALE.cov), sample.nobs = c(5312,6008))
summary(NELS.twogroup.fit, fit.measures = TRUE, standardized = TRUE, rsquare = TRUE)## lavaan 0.6-8 ended normally after 55 iterations
##
## Estimator ML
## Optimization method NLMINB
## Number of model parameters 36
##
## Number of observations per group:
## Group 1 5312
## Group 2 6008
##
## Model Test User Model:
##
## Test statistic 345.277
## Degrees of freedom 36
## P-value (Chi-square) 0.000
## Test statistic for each group:
## Group 1 146.970
## Group 2 198.307
##
## Model Test Baseline Model:
##
## Test statistic 16850.155
## Degrees of freedom 56
## P-value 0.000
##
## User Model versus Baseline Model:
##
## Comparative Fit Index (CFI) 0.982
## Tucker-Lewis Index (TLI) 0.971
##
## Loglikelihood and Information Criteria:
##
## Loglikelihood user model (H0) -87844.056
## Loglikelihood unrestricted model (H1) -87671.417
##
## Akaike (AIC) 175760.111
## Bayesian (BIC) 176024.147
## Sample-size adjusted Bayesian (BIC) 175909.743
##
## Root Mean Square Error of Approximation:
##
## RMSEA 0.039
## 90 Percent confidence interval - lower 0.035
## 90 Percent confidence interval - upper 0.043
## P-value RMSEA <= 0.05 1.000
##
## Standardized Root Mean Square Residual:
##
## SRMR 0.023
##
## Parameter Estimates:
##
## Standard errors Standard
## Information Expected
## Information saturated (h1) model Structured
##
##
## Group 1 [Group 1]:
##
## Latent Variables:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## f1 =~
## byconf1 1.000 0.379 0.647
## byconf2 0.863 0.031 27.486 0.000 0.327 0.509
## byconf3 0.701 0.029 23.820 0.000 0.266 0.428
## byconf4 1.272 0.040 31.424 0.000 0.482 0.738
## f2 =~
## byloc1 1.000 0.412 0.512
## byloc2 0.667 0.036 18.716 0.000 0.274 0.367
## byloc3 1.115 0.045 24.711 0.000 0.459 0.602
## byloc4 1.179 0.047 24.971 0.000 0.485 0.634
##
## Covariances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .byconf2 ~~
## .byconf3 0.079 0.005 15.304 0.000 0.079 0.255
## f1 ~~
## f2 0.085 0.004 18.978 0.000 0.546 0.546
##
## Variances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .byconf1 0.200 0.006 35.342 0.000 0.200 0.582
## .byconf2 0.306 0.007 43.856 0.000 0.306 0.741
## .byconf3 0.314 0.007 46.432 0.000 0.314 0.817
## .byconf4 0.194 0.007 25.998 0.000 0.194 0.455
## .byloc1 0.477 0.011 42.099 0.000 0.477 0.738
## .byloc2 0.484 0.010 47.493 0.000 0.484 0.865
## .byloc3 0.371 0.010 36.188 0.000 0.371 0.638
## .byloc4 0.351 0.010 33.480 0.000 0.351 0.599
## f1 0.144 0.007 21.119 0.000 1.000 1.000
## f2 0.169 0.011 15.846 0.000 1.000 1.000
##
## R-Square:
## Estimate
## byconf1 0.418
## byconf2 0.259
## byconf3 0.183
## byconf4 0.545
## byloc1 0.262
## byloc2 0.135
## byloc3 0.362
## byloc4 0.401
##
##
## Group 2 [Group 2]:
##
## Latent Variables:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## f1 =~
## byconf1 1.000 0.445 0.722
## byconf2 0.820 0.022 36.510 0.000 0.365 0.561
## byconf3 0.677 0.022 30.855 0.000 0.301 0.469
## byconf4 1.240 0.028 43.702 0.000 0.552 0.796
## f2 =~
## byloc1 1.000 0.441 0.548
## byloc2 0.604 0.027 22.120 0.000 0.267 0.379
## byloc3 1.091 0.035 30.827 0.000 0.482 0.639
## byloc4 1.261 0.040 31.500 0.000 0.556 0.699
##
## Covariances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .byconf2 ~~
## .byconf3 0.073 0.005 15.686 0.000 0.073 0.240
## f1 ~~
## f2 0.112 0.005 23.313 0.000 0.571 0.571
##
## Variances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .byconf1 0.181 0.005 36.091 0.000 0.181 0.478
## .byconf2 0.289 0.006 47.312 0.000 0.289 0.685
## .byconf3 0.321 0.006 50.052 0.000 0.321 0.780
## .byconf4 0.176 0.007 26.934 0.000 0.176 0.366
## .byloc1 0.453 0.010 45.301 0.000 0.453 0.700
## .byloc2 0.423 0.008 51.184 0.000 0.423 0.856
## .byloc3 0.336 0.009 39.152 0.000 0.336 0.592
## .byloc4 0.324 0.010 33.420 0.000 0.324 0.512
## f1 0.198 0.007 27.642 0.000 1.000 1.000
## f2 0.195 0.010 19.176 0.000 1.000 1.000
##
## R-Square:
## Estimate
## byconf1 0.522
## byconf2 0.315
## byconf3 0.220
## byconf4 0.634
## byloc1 0.300
## byloc2 0.144
## byloc3 0.408
## byloc4 0.4886.1.2.2 constrain factor loadings (metric invariance)
NELS.twogroup.fit.metric <- cfa(NELS.cfa.model, sample.cov = list(NELS.MALE.cov, NELS.FEMALE.cov), sample.nobs = c(5312,6008), group.equal = "loadings")
summary(NELS.twogroup.fit.metric, fit.measures = TRUE, standardized = TRUE, rsquare = TRUE, modindices = TRUE)## lavaan 0.6-8 ended normally after 38 iterations
##
## Estimator ML
## Optimization method NLMINB
## Number of model parameters 36
## Number of equality constraints 6
##
## Number of observations per group:
## Group 1 5312
## Group 2 6008
##
## Model Test User Model:
##
## Test statistic 354.073
## Degrees of freedom 42
## P-value (Chi-square) 0.000
## Test statistic for each group:
## Group 1 152.399
## Group 2 201.674
##
## Model Test Baseline Model:
##
## Test statistic 16850.155
## Degrees of freedom 56
## P-value 0.000
##
## User Model versus Baseline Model:
##
## Comparative Fit Index (CFI) 0.981
## Tucker-Lewis Index (TLI) 0.975
##
## Loglikelihood and Information Criteria:
##
## Loglikelihood user model (H0) -87848.453
## Loglikelihood unrestricted model (H1) -87671.417
##
## Akaike (AIC) 175756.907
## Bayesian (BIC) 175976.936
## Sample-size adjusted Bayesian (BIC) 175881.600
##
## Root Mean Square Error of Approximation:
##
## RMSEA 0.036
## 90 Percent confidence interval - lower 0.033
## 90 Percent confidence interval - upper 0.040
## P-value RMSEA <= 0.05 1.000
##
## Standardized Root Mean Square Residual:
##
## SRMR 0.024
##
## Parameter Estimates:
##
## Standard errors Standard
## Information Expected
## Information saturated (h1) model Structured
##
##
## Group 1 [Group 1]:
##
## Latent Variables:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## f1 =~
## byconf1 1.000 0.384 0.654
## byconf2 (.p2.) 0.835 0.018 45.707 0.000 0.321 0.501
## byconf3 (.p3.) 0.685 0.018 39.002 0.000 0.263 0.425
## byconf4 (.p4.) 1.251 0.023 53.805 0.000 0.481 0.736
## f2 =~
## byloc1 1.000 0.409 0.509
## byloc2 (.p6.) 0.628 0.022 28.972 0.000 0.257 0.345
## byloc3 (.p7.) 1.101 0.028 39.507 0.000 0.450 0.592
## byloc4 (.p8.) 1.228 0.031 40.163 0.000 0.502 0.652
##
## Covariances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .byconf2 ~~
## .byconf3 0.080 0.005 16.041 0.000 0.080 0.257
## f1 ~~
## f2 0.086 0.004 22.131 0.000 0.545 0.545
##
## Variances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .byconf1 0.198 0.005 38.071 0.000 0.198 0.572
## .byconf2 0.308 0.007 45.649 0.000 0.308 0.749
## .byconf3 0.314 0.007 47.520 0.000 0.314 0.819
## .byconf4 0.195 0.006 30.236 0.000 0.195 0.458
## .byloc1 0.479 0.011 44.102 0.000 0.479 0.741
## .byloc2 0.488 0.010 48.755 0.000 0.488 0.881
## .byloc3 0.375 0.009 39.720 0.000 0.375 0.649
## .byloc4 0.342 0.010 35.160 0.000 0.342 0.575
## f1 0.148 0.005 27.846 0.000 1.000 1.000
## f2 0.167 0.008 21.458 0.000 1.000 1.000
##
## R-Square:
## Estimate
## byconf1 0.428
## byconf2 0.251
## byconf3 0.181
## byconf4 0.542
## byloc1 0.259
## byloc2 0.119
## byloc3 0.351
## byloc4 0.425
##
##
## Group 2 [Group 2]:
##
## Latent Variables:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## f1 =~
## byconf1 1.000 0.442 0.719
## byconf2 (.p2.) 0.835 0.018 45.707 0.000 0.369 0.566
## byconf3 (.p3.) 0.685 0.018 39.002 0.000 0.303 0.471
## byconf4 (.p4.) 1.251 0.023 53.805 0.000 0.552 0.797
## f2 =~
## byloc1 1.000 0.443 0.550
## byloc2 (.p6.) 0.628 0.022 28.972 0.000 0.278 0.394
## byloc3 (.p7.) 1.101 0.028 39.507 0.000 0.488 0.645
## byloc4 (.p8.) 1.228 0.031 40.163 0.000 0.544 0.687
##
## Covariances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .byconf2 ~~
## .byconf3 0.073 0.005 15.777 0.000 0.073 0.239
## f1 ~~
## f2 0.112 0.004 25.255 0.000 0.571 0.571
##
## Variances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .byconf1 0.183 0.005 38.036 0.000 0.183 0.483
## .byconf2 0.289 0.006 47.749 0.000 0.289 0.680
## .byconf3 0.321 0.006 50.418 0.000 0.321 0.778
## .byconf4 0.175 0.006 28.604 0.000 0.175 0.365
## .byloc1 0.452 0.010 46.178 0.000 0.452 0.697
## .byloc2 0.421 0.008 51.230 0.000 0.421 0.845
## .byloc3 0.334 0.008 40.208 0.000 0.334 0.584
## .byloc4 0.331 0.009 36.474 0.000 0.331 0.528
## f1 0.195 0.006 30.645 0.000 1.000 1.000
## f2 0.196 0.009 22.733 0.000 1.000 1.000
##
## R-Square:
## Estimate
## byconf1 0.517
## byconf2 0.320
## byconf3 0.222
## byconf4 0.635
## byloc1 0.303
## byloc2 0.155
## byloc3 0.416
## byloc4 0.472
##
## Modification Indices:
##
## lhs op rhs block group level mi epc sepc.lv sepc.all
## 1 f1 =~ byconf1 1 1 1 0.992 -0.035 -0.013 -0.023
## 5 f2 =~ byloc1 1 1 1 0.001 -0.002 -0.001 -0.001
## 21 f1 =~ byconf1 2 2 1 0.992 0.035 0.015 0.025
## 25 f2 =~ byloc1 2 2 1 0.001 0.002 0.001 0.001
## 47 f1 =~ byloc1 1 1 1 7.415 0.099 0.038 0.048
## 48 f1 =~ byloc2 1 1 1 1.455 -0.041 -0.016 -0.021
## 49 f1 =~ byloc3 1 1 1 0.171 -0.015 -0.006 -0.007
## 50 f1 =~ byloc4 1 1 1 1.388 -0.043 -0.017 -0.022
## 51 f2 =~ byconf1 1 1 1 5.214 -0.057 -0.023 -0.040
## 52 f2 =~ byconf2 1 1 1 1.531 0.032 0.013 0.020
## 53 f2 =~ byconf3 1 1 1 1.644 0.032 0.013 0.021
## 54 f2 =~ byconf4 1 1 1 0.150 0.011 0.005 0.007
## 55 byconf1 ~~ byconf2 1 1 1 0.103 0.001 0.001 0.005
## 56 byconf1 ~~ byconf3 1 1 1 0.042 -0.001 -0.001 -0.003
## 57 byconf1 ~~ byconf4 1 1 1 0.546 0.005 0.005 0.023
## 58 byconf1 ~~ byloc1 1 1 1 3.609 0.010 0.010 0.032
## 59 byconf1 ~~ byloc2 1 1 1 40.513 -0.031 -0.031 -0.101
## 60 byconf1 ~~ byloc3 1 1 1 0.896 0.005 0.005 0.017
## 61 byconf1 ~~ byloc4 1 1 1 2.638 -0.008 -0.008 -0.030
## 62 byconf2 ~~ byconf4 1 1 1 0.056 -0.001 -0.001 -0.005
## 63 byconf2 ~~ byloc1 1 1 1 0.224 0.003 0.003 0.007
## 64 byconf2 ~~ byloc2 1 1 1 40.886 0.035 0.035 0.090
## 65 byconf2 ~~ byloc3 1 1 1 7.154 -0.014 -0.014 -0.041
## 66 byconf2 ~~ byloc4 1 1 1 0.013 0.001 0.001 0.002
## 67 byconf3 ~~ byconf4 1 1 1 0.466 -0.003 -0.003 -0.013
## 68 byconf3 ~~ byloc1 1 1 1 0.261 0.003 0.003 0.007
## 69 byconf3 ~~ byloc2 1 1 1 0.043 -0.001 -0.001 -0.003
## 70 byconf3 ~~ byloc3 1 1 1 0.012 -0.001 -0.001 -0.002
## 71 byconf3 ~~ byloc4 1 1 1 2.491 0.008 0.008 0.025
## 72 byconf4 ~~ byloc1 1 1 1 0.954 0.005 0.005 0.018
## 73 byconf4 ~~ byloc2 1 1 1 1.090 -0.006 -0.006 -0.018
## 74 byconf4 ~~ byloc3 1 1 1 0.001 0.000 0.000 0.001
## 75 byconf4 ~~ byloc4 1 1 1 0.021 0.001 0.001 0.003
## 76 byloc1 ~~ byloc2 1 1 1 37.675 0.046 0.046 0.095
## 77 byloc1 ~~ byloc3 1 1 1 0.019 -0.001 -0.001 -0.003
## 78 byloc1 ~~ byloc4 1 1 1 35.121 -0.050 -0.050 -0.122
## 79 byloc2 ~~ byloc3 1 1 1 4.045 -0.014 -0.014 -0.033
## 80 byloc2 ~~ byloc4 1 1 1 0.219 0.003 0.003 0.008
## 81 byloc3 ~~ byloc4 1 1 1 11.328 0.029 0.029 0.082
## 82 f1 =~ byloc1 2 2 1 7.771 0.084 0.037 0.046
## 83 f1 =~ byloc2 2 2 1 16.414 -0.109 -0.048 -0.068
## 84 f1 =~ byloc3 2 2 1 8.436 -0.084 -0.037 -0.049
## 85 f1 =~ byloc4 2 2 1 8.086 0.088 0.039 0.049
## 86 f2 =~ byconf1 2 2 1 24.080 -0.111 -0.049 -0.080
## 87 f2 =~ byconf2 2 2 1 2.203 0.034 0.015 0.023
## 88 f2 =~ byconf3 2 2 1 5.679 0.055 0.025 0.038
## 89 f2 =~ byconf4 2 2 1 3.915 0.053 0.024 0.034
## 90 byconf1 ~~ byconf2 2 2 1 5.722 0.010 0.010 0.042
## 91 byconf1 ~~ byconf3 2 2 1 0.088 -0.001 -0.001 -0.005
## 92 byconf1 ~~ byconf4 2 2 1 15.224 0.027 0.027 0.149
## 93 byconf1 ~~ byloc1 2 2 1 0.283 -0.002 -0.002 -0.008
## 94 byconf1 ~~ byloc2 2 2 1 57.623 -0.032 -0.032 -0.115
## 95 byconf1 ~~ byloc3 2 2 1 0.703 -0.003 -0.003 -0.014
## 96 byconf1 ~~ byloc4 2 2 1 1.048 -0.004 -0.004 -0.018
## 97 byconf2 ~~ byconf4 2 2 1 16.943 -0.020 -0.020 -0.088
## 98 byconf2 ~~ byloc1 2 2 1 2.122 0.007 0.007 0.020
## 99 byconf2 ~~ byloc2 2 2 1 18.095 0.020 0.020 0.057
## 100 byconf2 ~~ byloc3 2 2 1 5.209 -0.010 -0.010 -0.033
## 101 byconf2 ~~ byloc4 2 2 1 1.252 0.005 0.005 0.017
## 102 byconf3 ~~ byconf4 2 2 1 1.919 -0.006 -0.006 -0.027
## 103 byconf3 ~~ byloc1 2 2 1 0.165 -0.002 -0.002 -0.006
## 104 byconf3 ~~ byloc2 2 2 1 1.191 0.005 0.005 0.014
## 105 byconf3 ~~ byloc3 2 2 1 3.918 0.009 0.009 0.028
## 106 byconf3 ~~ byloc4 2 2 1 0.992 0.005 0.005 0.015
## 107 byconf4 ~~ byloc1 2 2 1 6.377 0.013 0.013 0.045
## 108 byconf4 ~~ byloc2 2 2 1 0.035 -0.001 -0.001 -0.003
## 109 byconf4 ~~ byloc3 2 2 1 1.394 -0.005 -0.005 -0.022
## 110 byconf4 ~~ byloc4 2 2 1 2.222 0.007 0.007 0.030
## 111 byloc1 ~~ byloc2 2 2 1 55.136 0.048 0.048 0.110
## 112 byloc1 ~~ byloc3 2 2 1 7.479 -0.020 -0.020 -0.051
## 113 byloc1 ~~ byloc4 2 2 1 20.270 -0.035 -0.035 -0.092
## 114 byloc2 ~~ byloc3 2 2 1 5.065 -0.014 -0.014 -0.036
## 115 byloc2 ~~ byloc4 2 2 1 5.493 -0.015 -0.015 -0.040
## 116 byloc3 ~~ byloc4 2 2 1 32.756 0.047 0.047 0.143
## sepc.nox
## 1 -0.023
## 5 -0.001
## 21 0.025
## 25 0.001
## 47 0.048
## 48 -0.021
## 49 -0.007
## 50 -0.022
## 51 -0.040
## 52 0.020
## 53 0.021
## 54 0.007
## 55 0.005
## 56 -0.003
## 57 0.023
## 58 0.032
## 59 -0.101
## 60 0.017
## 61 -0.030
## 62 -0.005
## 63 0.007
## 64 0.090
## 65 -0.041
## 66 0.002
## 67 -0.013
## 68 0.007
## 69 -0.003
## 70 -0.002
## 71 0.025
## 72 0.018
## 73 -0.018
## 74 0.001
## 75 0.003
## 76 0.095
## 77 -0.003
## 78 -0.122
## 79 -0.033
## 80 0.008
## 81 0.082
## 82 0.046
## 83 -0.068
## 84 -0.049
## 85 0.049
## 86 -0.080
## 87 0.023
## 88 0.038
## 89 0.034
## 90 0.042
## 91 -0.005
## 92 0.149
## 93 -0.008
## 94 -0.115
## 95 -0.014
## 96 -0.018
## 97 -0.088
## 98 0.020
## 99 0.057
## 100 -0.033
## 101 0.017
## 102 -0.027
## 103 -0.006
## 104 0.014
## 105 0.028
## 106 0.015
## 107 0.045
## 108 -0.003
## 109 -0.022
## 110 0.030
## 111 0.110
## 112 -0.051
## 113 -0.092
## 114 -0.036
## 115 -0.040
## 116 0.143Plot the path diagram
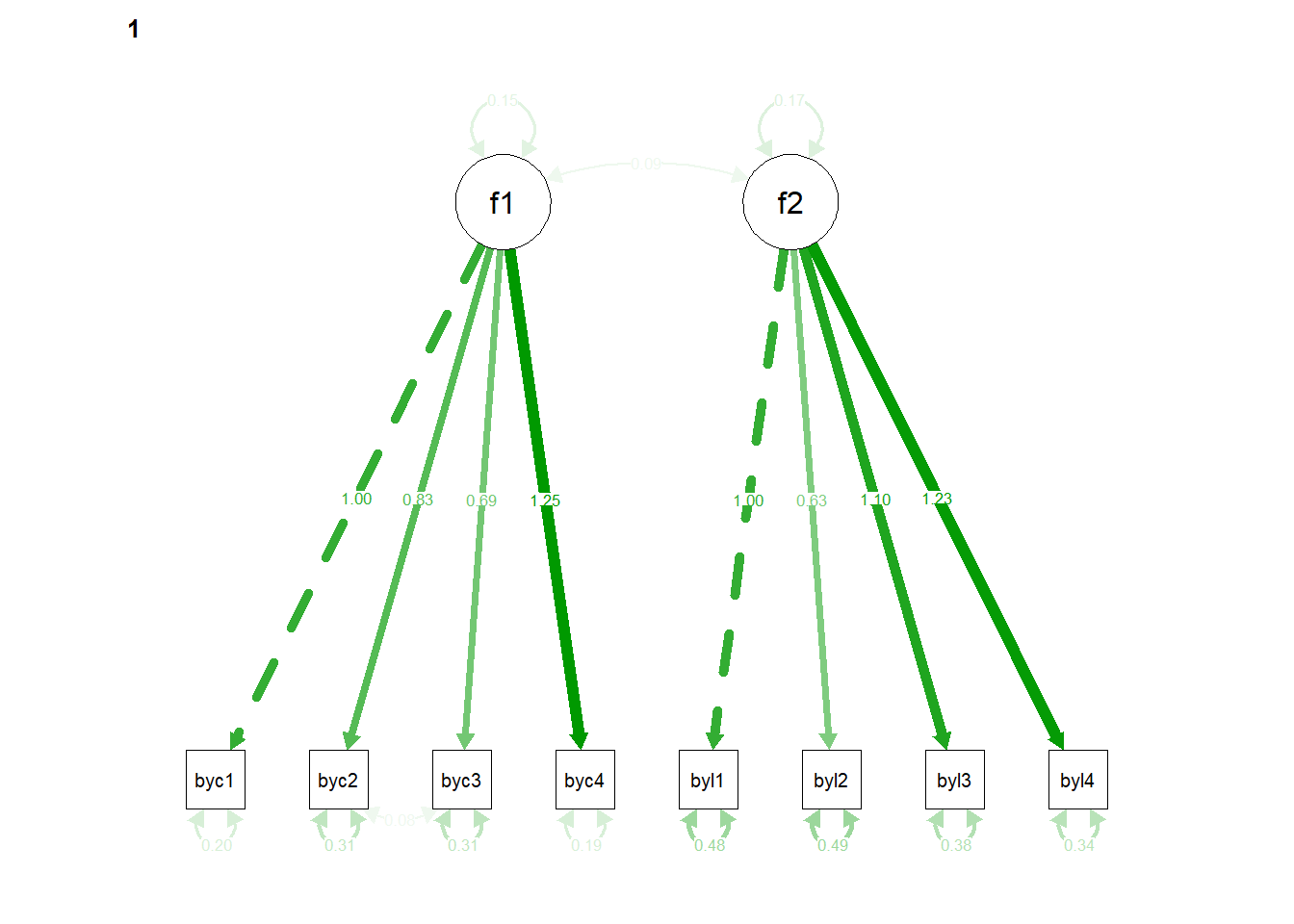
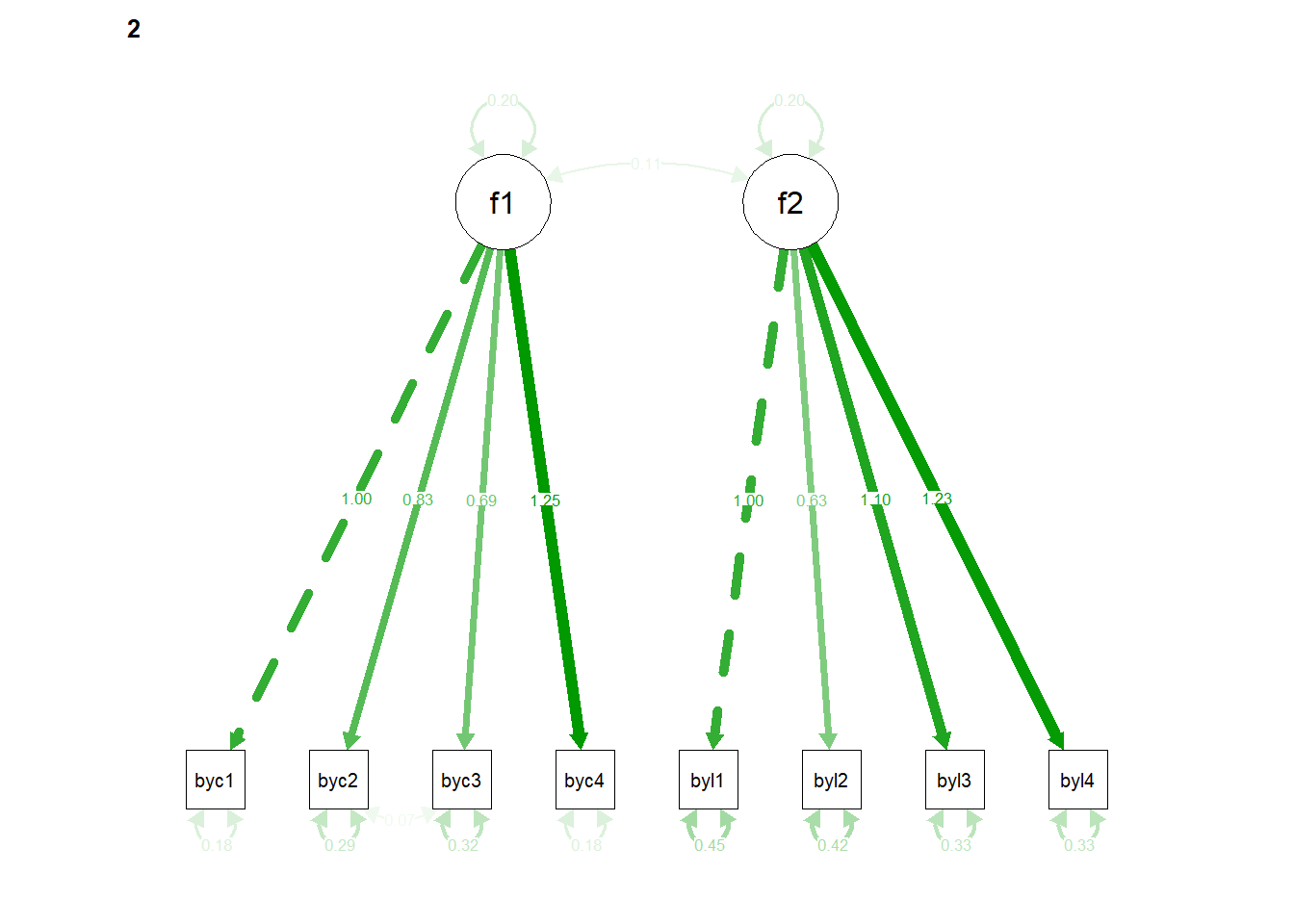
6.1.2.3 compare nested models
## Chi-Squared Difference Test
##
## Df AIC BIC Chisq Chisq diff Df diff Pr(>Chisq)
## NELS.twogroup.fit 36 175760 176024 345.28
## NELS.twogroup.fit.metric 42 175757 175977 354.07 8.7955 6 0.1854## Chi-Squared Difference Test
##
## Df AIC BIC Chisq Chisq diff Df diff Pr(>Chisq)
## NELS.twogroup.fit 36 175760 176024 345.28
## NELS.twogroup.fit.metric 42 175757 175977 354.07 8.7955 6 0.18546.1.2.4 use modindices() function to release constraints across groups
By default, the modindices() function will only consider nonfree parameters (those fixed at 0 or 1). Parameters that are constrained to be equal across groups are still free parameters. Set the argument “free.remove = FALSE” for releasing constraints across groups.
mi <- modindices(NELS.twogroup.fit.metric, free.remove = FALSE)
mi[mi$op == "=~",] #request modification indices for factor loadings only## lhs op rhs block group level mi epc sepc.lv sepc.all sepc.nox
## 1 f1 =~ byconf1 1 1 1 0.992 -0.035 -0.013 -0.023 -0.023
## 2 f1 =~ byconf2 1 1 1 0.365 0.014 0.005 0.008 0.008
## 3 f1 =~ byconf3 1 1 1 0.001 0.001 0.000 0.001 0.001
## 4 f1 =~ byconf4 1 1 1 0.008 0.002 0.001 0.001 0.001
## 5 f2 =~ byloc1 1 1 1 0.001 -0.002 -0.001 -0.001 -0.001
## 6 f2 =~ byloc2 1 1 1 1.939 0.040 0.016 0.022 0.022
## 7 f2 =~ byloc3 1 1 1 0.460 0.018 0.007 0.010 0.010
## 8 f2 =~ byloc4 1 1 1 1.453 -0.031 -0.013 -0.016 -0.016
## 21 f1 =~ byconf1 2 2 1 0.992 0.035 0.015 0.025 0.025
## 22 f1 =~ byconf2 2 2 1 0.215 -0.008 -0.004 -0.006 -0.006
## 23 f1 =~ byconf3 2 2 1 0.001 -0.001 0.000 0.000 0.000
## 24 f1 =~ byconf4 2 2 1 0.005 -0.001 -0.001 -0.001 -0.001
## 25 f2 =~ byloc1 2 2 1 0.001 0.002 0.001 0.001 0.001
## 26 f2 =~ byloc2 2 2 1 1.185 -0.025 -0.011 -0.015 -0.015
## 27 f2 =~ byloc3 2 2 1 0.298 -0.012 -0.005 -0.007 -0.007
## 28 f2 =~ byloc4 2 2 1 1.022 0.022 0.010 0.012 0.012
## 47 f1 =~ byloc1 1 1 1 7.415 0.099 0.038 0.048 0.048
## 48 f1 =~ byloc2 1 1 1 1.455 -0.041 -0.016 -0.021 -0.021
## 49 f1 =~ byloc3 1 1 1 0.171 -0.015 -0.006 -0.007 -0.007
## 50 f1 =~ byloc4 1 1 1 1.388 -0.043 -0.017 -0.022 -0.022
## 51 f2 =~ byconf1 1 1 1 5.214 -0.057 -0.023 -0.040 -0.040
## 52 f2 =~ byconf2 1 1 1 1.531 0.032 0.013 0.020 0.020
## 53 f2 =~ byconf3 1 1 1 1.644 0.032 0.013 0.021 0.021
## 54 f2 =~ byconf4 1 1 1 0.150 0.011 0.005 0.007 0.007
## 82 f1 =~ byloc1 2 2 1 7.771 0.084 0.037 0.046 0.046
## 83 f1 =~ byloc2 2 2 1 16.414 -0.109 -0.048 -0.068 -0.068
## 84 f1 =~ byloc3 2 2 1 8.436 -0.084 -0.037 -0.049 -0.049
## 85 f1 =~ byloc4 2 2 1 8.086 0.088 0.039 0.049 0.049
## 86 f2 =~ byconf1 2 2 1 24.080 -0.111 -0.049 -0.080 -0.080
## 87 f2 =~ byconf2 2 2 1 2.203 0.034 0.015 0.023 0.023
## 88 f2 =~ byconf3 2 2 1 5.679 0.055 0.025 0.038 0.038
## 89 f2 =~ byconf4 2 2 1 3.915 0.053 0.024 0.034 0.034NELS.twogroup.fit.metric.partial <- cfa(NELS.cfa.model, sample.cov = list(NELS.MALE.cov, NELS.FEMALE.cov), sample.nobs = c(5312,6008), group.equal = "loadings", group.partial = c("f2=~byloc2")) #release one factor loading constraint
summary(NELS.twogroup.fit.metric.partial, fit.measures = TRUE, standardized = TRUE, rsquare = TRUE,)## lavaan 0.6-8 ended normally after 43 iterations
##
## Estimator ML
## Optimization method NLMINB
## Number of model parameters 36
## Number of equality constraints 5
##
## Number of observations per group:
## Group 1 5312
## Group 2 6008
##
## Model Test User Model:
##
## Test statistic 350.532
## Degrees of freedom 41
## P-value (Chi-square) 0.000
## Test statistic for each group:
## Group 1 150.269
## Group 2 200.264
##
## Model Test Baseline Model:
##
## Test statistic 16850.155
## Degrees of freedom 56
## P-value 0.000
##
## User Model versus Baseline Model:
##
## Comparative Fit Index (CFI) 0.982
## Tucker-Lewis Index (TLI) 0.975
##
## Loglikelihood and Information Criteria:
##
## Loglikelihood user model (H0) -87846.683
## Loglikelihood unrestricted model (H1) -87671.417
##
## Akaike (AIC) 175755.366
## Bayesian (BIC) 175982.731
## Sample-size adjusted Bayesian (BIC) 175884.216
##
## Root Mean Square Error of Approximation:
##
## RMSEA 0.037
## 90 Percent confidence interval - lower 0.033
## 90 Percent confidence interval - upper 0.040
## P-value RMSEA <= 0.05 1.000
##
## Standardized Root Mean Square Residual:
##
## SRMR 0.024
##
## Parameter Estimates:
##
## Standard errors Standard
## Information Expected
## Information saturated (h1) model Structured
##
##
## Group 1 [Group 1]:
##
## Latent Variables:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## f1 =~
## byconf1 1.000 0.384 0.654
## byconf2 (.p2.) 0.835 0.018 45.711 0.000 0.321 0.501
## byconf3 (.p3.) 0.685 0.018 39.002 0.000 0.263 0.425
## byconf4 (.p4.) 1.251 0.023 53.806 0.000 0.481 0.736
## f2 =~
## byloc1 1.000 0.406 0.506
## byloc2 0.675 0.033 20.264 0.000 0.274 0.367
## byloc3 (.p7.) 1.100 0.028 39.509 0.000 0.447 0.589
## byloc4 (.p8.) 1.229 0.031 40.182 0.000 0.499 0.649
##
## Covariances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .byconf2 ~~
## .byconf3 0.080 0.005 16.033 0.000 0.080 0.257
## f1 ~~
## f2 0.085 0.004 22.058 0.000 0.545 0.545
##
## Variances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .byconf1 0.198 0.005 38.074 0.000 0.198 0.573
## .byconf2 0.308 0.007 45.644 0.000 0.308 0.749
## .byconf3 0.314 0.007 47.518 0.000 0.314 0.819
## .byconf4 0.195 0.006 30.233 0.000 0.195 0.458
## .byloc1 0.479 0.011 44.167 0.000 0.479 0.744
## .byloc2 0.484 0.010 47.514 0.000 0.484 0.866
## .byloc3 0.377 0.009 39.855 0.000 0.377 0.653
## .byloc4 0.343 0.010 35.250 0.000 0.343 0.579
## f1 0.148 0.005 27.845 0.000 1.000 1.000
## f2 0.165 0.008 21.186 0.000 1.000 1.000
##
## R-Square:
## Estimate
## byconf1 0.427
## byconf2 0.251
## byconf3 0.181
## byconf4 0.542
## byloc1 0.256
## byloc2 0.134
## byloc3 0.347
## byloc4 0.421
##
##
## Group 2 [Group 2]:
##
## Latent Variables:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## f1 =~
## byconf1 1.000 0.442 0.719
## byconf2 (.p2.) 0.835 0.018 45.711 0.000 0.369 0.566
## byconf3 (.p3.) 0.685 0.018 39.002 0.000 0.303 0.471
## byconf4 (.p4.) 1.251 0.023 53.806 0.000 0.552 0.797
## f2 =~
## byloc1 1.000 0.445 0.552
## byloc2 0.601 0.026 23.218 0.000 0.267 0.380
## byloc3 (.p7.) 1.100 0.028 39.509 0.000 0.489 0.647
## byloc4 (.p8.) 1.229 0.031 40.182 0.000 0.546 0.689
##
## Covariances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .byconf2 ~~
## .byconf3 0.073 0.005 15.779 0.000 0.073 0.239
## f1 ~~
## f2 0.112 0.004 25.240 0.000 0.572 0.572
##
## Variances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .byconf1 0.183 0.005 38.038 0.000 0.183 0.483
## .byconf2 0.289 0.006 47.749 0.000 0.289 0.680
## .byconf3 0.321 0.006 50.418 0.000 0.321 0.778
## .byconf4 0.175 0.006 28.612 0.000 0.175 0.365
## .byloc1 0.452 0.010 46.096 0.000 0.452 0.696
## .byloc2 0.423 0.008 51.157 0.000 0.423 0.856
## .byloc3 0.333 0.008 40.035 0.000 0.333 0.582
## .byloc4 0.330 0.009 36.170 0.000 0.330 0.525
## f1 0.195 0.006 30.645 0.000 1.000 1.000
## f2 0.198 0.009 22.647 0.000 1.000 1.000
##
## R-Square:
## Estimate
## byconf1 0.517
## byconf2 0.320
## byconf3 0.222
## byconf4 0.635
## byloc1 0.304
## byloc2 0.144
## byloc3 0.418
## byloc4 0.4756.1.2.5 constrain loadings, residual variances, and residual covariances
The value for “group.equal” should be a vector of character strings. They can be one or more of the following: “loadings”, “intercepts”, “means”, “thresholds”, “regressions”, “residuals”, “residual.covariances”, “lv.variances” or “lv.covariances”, specifying the pattern of equality constraints across multiple groups.
NELS.twogroup.fit.metric.constrain.residuals.residualcovariances <- cfa(NELS.cfa.model, sample.cov = list(NELS.MALE.cov, NELS.FEMALE.cov), sample.nobs = c(5312,6008), group.equal = c("loadings", "residuals", "residual.covariances"))
summary(NELS.twogroup.fit.metric.constrain.residuals.residualcovariances, fit.measures = TRUE, standardized = TRUE, rsquare = TRUE)## lavaan 0.6-8 ended normally after 35 iterations
##
## Estimator ML
## Optimization method NLMINB
## Number of model parameters 36
## Number of equality constraints 15
##
## Number of observations per group:
## Group 1 5312
## Group 2 6008
##
## Model Test User Model:
##
## Test statistic 419.715
## Degrees of freedom 51
## P-value (Chi-square) 0.000
## Test statistic for each group:
## Group 1 185.577
## Group 2 234.138
##
## Model Test Baseline Model:
##
## Test statistic 16850.155
## Degrees of freedom 56
## P-value 0.000
##
## User Model versus Baseline Model:
##
## Comparative Fit Index (CFI) 0.978
## Tucker-Lewis Index (TLI) 0.976
##
## Loglikelihood and Information Criteria:
##
## Loglikelihood user model (H0) -87881.275
## Loglikelihood unrestricted model (H1) -87671.417
##
## Akaike (AIC) 175804.549
## Bayesian (BIC) 175958.570
## Sample-size adjusted Bayesian (BIC) 175891.835
##
## Root Mean Square Error of Approximation:
##
## RMSEA 0.036
## 90 Percent confidence interval - lower 0.033
## 90 Percent confidence interval - upper 0.039
## P-value RMSEA <= 0.05 1.000
##
## Standardized Root Mean Square Residual:
##
## SRMR 0.028
##
## Parameter Estimates:
##
## Standard errors Standard
## Information Expected
## Information saturated (h1) model Structured
##
##
## Group 1 [Group 1]:
##
## Latent Variables:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## f1 =~
## byconf1 1.000 0.388 0.664
## byconf2 (.p2.) 0.837 0.018 45.507 0.000 0.324 0.511
## byconf3 (.p3.) 0.687 0.018 38.969 0.000 0.266 0.427
## byconf4 (.p4.) 1.252 0.023 53.506 0.000 0.485 0.749
## f2 =~
## byloc1 1.000 0.413 0.519
## byloc2 (.p6.) 0.631 0.022 28.886 0.000 0.261 0.362
## byloc3 (.p7.) 1.101 0.028 39.370 0.000 0.455 0.608
## byloc4 (.p8.) 1.227 0.031 40.050 0.000 0.507 0.658
##
## Covariances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .byconf2 ~~
## .byconf3 (.p9.) 0.076 0.003 21.938 0.000 0.076 0.247
## f1 ~~
## f2 0.086 0.004 22.086 0.000 0.534 0.534
##
## Variances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .byconf1 (.10.) 0.190 0.004 50.559 0.000 0.190 0.558
## .byconf2 (.11.) 0.297 0.005 64.604 0.000 0.297 0.739
## .byconf3 (.12.) 0.318 0.005 68.309 0.000 0.318 0.818
## .byconf4 (.13.) 0.185 0.005 37.663 0.000 0.185 0.440
## .byloc1 (.14.) 0.464 0.008 61.889 0.000 0.464 0.731
## .byloc2 (.15.) 0.452 0.006 69.908 0.000 0.452 0.869
## .byloc3 (.16.) 0.353 0.007 53.462 0.000 0.353 0.630
## .byloc4 (.17.) 0.336 0.007 47.134 0.000 0.336 0.567
## f1 0.150 0.005 28.430 0.000 1.000 1.000
## f2 0.171 0.008 21.772 0.000 1.000 1.000
##
## R-Square:
## Estimate
## byconf1 0.442
## byconf2 0.261
## byconf3 0.182
## byconf4 0.560
## byloc1 0.269
## byloc2 0.131
## byloc3 0.370
## byloc4 0.433
##
##
## Group 2 [Group 2]:
##
## Latent Variables:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## f1 =~
## byconf1 1.000 0.439 0.709
## byconf2 (.p2.) 0.837 0.018 45.507 0.000 0.367 0.558
## byconf3 (.p3.) 0.687 0.018 38.969 0.000 0.301 0.471
## byconf4 (.p4.) 1.252 0.023 53.506 0.000 0.549 0.787
## f2 =~
## byloc1 1.000 0.439 0.542
## byloc2 (.p6.) 0.631 0.022 28.886 0.000 0.277 0.381
## byloc3 (.p7.) 1.101 0.028 39.370 0.000 0.484 0.631
## byloc4 (.p8.) 1.227 0.031 40.050 0.000 0.539 0.681
##
## Covariances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .byconf2 ~~
## .byconf3 (.p9.) 0.076 0.003 21.938 0.000 0.076 0.247
## f1 ~~
## f2 0.112 0.004 25.265 0.000 0.581 0.581
##
## Variances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .byconf1 (.10.) 0.190 0.004 50.559 0.000 0.190 0.497
## .byconf2 (.11.) 0.297 0.005 64.604 0.000 0.297 0.688
## .byconf3 (.12.) 0.318 0.005 68.309 0.000 0.318 0.778
## .byconf4 (.13.) 0.185 0.005 37.663 0.000 0.185 0.380
## .byloc1 (.14.) 0.464 0.008 61.889 0.000 0.464 0.706
## .byloc2 (.15.) 0.452 0.006 69.908 0.000 0.452 0.855
## .byloc3 (.16.) 0.353 0.007 53.462 0.000 0.353 0.601
## .byloc4 (.17.) 0.336 0.007 47.134 0.000 0.336 0.537
## f1 0.192 0.006 30.451 0.000 1.000 1.000
## f2 0.193 0.009 22.666 0.000 1.000 1.000
##
## R-Square:
## Estimate
## byconf1 0.503
## byconf2 0.312
## byconf3 0.222
## byconf4 0.620
## byloc1 0.294
## byloc2 0.145
## byloc3 0.399
## byloc4 0.4636.1.3 CFA with Ordered Categorical Variables – An example from Wang & Su (2013)
NELS.SelfConcept.3waves.dat <- read.fwf("data/NELSSelfConcept3waves.dat",
widths=c(7,4,2,2,11,rep(2,39)),header = FALSE,
col.names = c("id", "status", "race", "gender", "weight", c(paste0('byconf', 1:7, sep=""), paste0('byloc', 1:6, sep=""), paste0('f1conf', 1:7, sep=""), paste0('f1loc', 1:6, sep=""), paste0('f2conf', 1:7, sep=""), paste0('f2loc', 1:6, sep=""))))
#recode 99 as NA for missing values
vars.99.as.missing <- names(NELS.SelfConcept.3waves.dat) %in% c("race", "gender", c(paste0('byconf', 1:7, sep=""), paste0('byloc', 1:6, sep=""),
paste0('f1conf', 1:7, sep=""), paste0('f1loc', 1:6, sep=""),
paste0('f2conf', 1:7, sep=""), paste0('f2loc', 1:6, sep="")))
dat.with.missing <- NELS.SelfConcept.3waves.dat[vars.99.as.missing]
dat.with.missing[dat.with.missing==99] <- NA
#add back variables that has no missing values. Three variables have no missing values: id, status, and weight.
NELS.SelfConcept.3waves.dat.NA.as.missing <- cbind(NELS.SelfConcept.3waves.dat[!vars.99.as.missing],dat.with.missing)
#calcuate total missing values in each column
colSums(is.na(NELS.SelfConcept.3waves.dat.NA.as.missing))## id status weight race gender byconf1 byconf2 byconf3 byconf4 byconf5
## 0 0 0 987 760 853 981 931 943 952
## byconf6 byconf7 byloc1 byloc2 byloc3 byloc4 byloc5 byloc6 f1conf1 f1conf2
## 945 918 875 901 896 893 917 890 1528 1573
## f1conf3 f1conf4 f1conf5 f1conf6 f1conf7 f1loc1 f1loc2 f1loc3 f1loc4 f1loc5
## 1621 1609 1633 1613 1634 1575 1591 1584 1625 1631
## f1loc6 f2conf1 f2conf2 f2conf3 f2conf4 f2conf5 f2conf6 f2conf7 f2loc1 f2loc2
## 1638 2279 2373 2373 2361 2356 2389 2380 2317 2331
## f2loc3 f2loc4 f2loc5 f2loc6
## 2347 2382 2381 2367NELS.SelfConcept.3waves.model <-
paste0('f1 =~ ', paste0('byconf', 1:4, collapse=' + '), ' \n',
' f2 =~ ', paste0('byloc', 1:4, collapse=' + '), ' \n',
' f3 =~ ', paste0('f1conf', 1:4, collapse=' + '), ' \n',
' f4 =~ ', paste0('f1loc', 1:4, collapse=' + '), ' \n',
' f5 =~ ', paste0('f2conf', 1:4, collapse=' + '), ' \n',
' f6 =~ ', paste0('f2loc', 1:4, collapse=' + '), ' \n')lavaan uses listwise deletion as the default. FIML can be enabled by specifying the argument missing=“ml” or missing=“fiml”. However, missing = “fiml” or “ml” is not available in the categorical setting. The default estimator for endogenous ordered categorical variables is WLSMV.
NELS.cfa.ordered.categorical.indicators.fit <- cfa(NELS.SelfConcept.3waves.model, data=NELS.SelfConcept.3waves.dat.NA.as.missing, ordered = c(paste0('byconf', 1:7, sep=""), paste0('byloc', 1:6, sep=""),
paste0('f1conf', 1:7, sep=""), paste0('f1loc', 1:6, sep=""), paste0('f2conf', 1:7, sep=""), paste0('f2loc', 1:6, sep="")))
summary(NELS.cfa.ordered.categorical.indicators.fit, fit.measures = TRUE, standardized = TRUE, rsquare = TRUE)## lavaan 0.6-8 ended normally after 54 iterations
##
## Estimator DWLS
## Optimization method NLMINB
## Number of model parameters 111
##
## Used Total
## Number of observations 7982 12144
##
## Model Test User Model:
## Standard Robust
## Test Statistic 4857.384 6358.478
## Degrees of freedom 237 237
## P-value (Chi-square) 0.000 0.000
## Scaling correction factor 0.770
## Shift parameter 53.452
## simple second-order correction
##
## Model Test Baseline Model:
##
## Test statistic 294010.961 131370.844
## Degrees of freedom 276 276
## P-value 0.000 0.000
## Scaling correction factor 2.241
##
## User Model versus Baseline Model:
##
## Comparative Fit Index (CFI) 0.984 0.953
## Tucker-Lewis Index (TLI) 0.982 0.946
##
## Robust Comparative Fit Index (CFI) NA
## Robust Tucker-Lewis Index (TLI) NA
##
## Root Mean Square Error of Approximation:
##
## RMSEA 0.049 0.057
## 90 Percent confidence interval - lower 0.048 0.056
## 90 Percent confidence interval - upper 0.051 0.058
## P-value RMSEA <= 0.05 0.781 0.000
##
## Robust RMSEA NA
## 90 Percent confidence interval - lower NA
## 90 Percent confidence interval - upper NA
##
## Standardized Root Mean Square Residual:
##
## SRMR 0.043 0.043
##
## Parameter Estimates:
##
## Standard errors Robust.sem
## Information Expected
## Information saturated (h1) model Unstructured
##
## Latent Variables:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## f1 =~
## byconf1 1.000 0.804 0.804
## byconf2 0.857 0.014 60.421 0.000 0.689 0.689
## byconf3 0.780 0.015 52.992 0.000 0.627 0.627
## byconf4 1.014 0.015 69.772 0.000 0.815 0.815
## f2 =~
## byloc1 1.000 0.608 0.608
## byloc2 0.731 0.026 28.172 0.000 0.445 0.445
## byloc3 1.088 0.027 40.686 0.000 0.662 0.662
## byloc4 1.191 0.028 41.935 0.000 0.725 0.725
## f3 =~
## f1conf1 1.000 0.816 0.816
## f1conf2 0.950 0.011 87.943 0.000 0.775 0.775
## f1conf3 0.885 0.011 78.145 0.000 0.722 0.722
## f1conf4 0.993 0.011 91.221 0.000 0.810 0.810
## f4 =~
## f1loc1 1.000 0.653 0.653
## f1loc2 0.750 0.019 38.514 0.000 0.490 0.490
## f1loc3 1.113 0.020 56.883 0.000 0.726 0.726
## f1loc4 1.194 0.021 57.859 0.000 0.780 0.780
## f5 =~
## f2conf1 1.000 0.832 0.832
## f2conf2 1.004 0.009 114.173 0.000 0.836 0.836
## f2conf3 0.966 0.009 107.139 0.000 0.804 0.804
## f2conf4 1.016 0.009 110.521 0.000 0.845 0.845
## f6 =~
## f2loc1 1.000 0.651 0.651
## f2loc2 0.825 0.019 42.435 0.000 0.537 0.537
## f2loc3 1.187 0.020 58.803 0.000 0.773 0.773
## f2loc4 1.247 0.021 59.160 0.000 0.812 0.812
##
## Covariances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## f1 ~~
## f2 0.282 0.008 33.859 0.000 0.577 0.577
## f3 0.380 0.009 42.865 0.000 0.579 0.579
## f4 0.183 0.008 23.303 0.000 0.349 0.349
## f5 0.314 0.009 35.663 0.000 0.470 0.470
## f6 0.145 0.008 18.486 0.000 0.278 0.278
## f2 ~~
## f3 0.180 0.008 22.709 0.000 0.363 0.363
## f4 0.236 0.008 31.124 0.000 0.594 0.594
## f5 0.154 0.008 19.334 0.000 0.305 0.305
## f6 0.194 0.007 27.343 0.000 0.490 0.490
## f3 ~~
## f4 0.329 0.008 43.147 0.000 0.618 0.618
## f5 0.426 0.008 51.411 0.000 0.627 0.627
## f6 0.206 0.008 26.909 0.000 0.388 0.388
## f4 ~~
## f5 0.219 0.008 27.761 0.000 0.404 0.404
## f6 0.266 0.007 35.853 0.000 0.627 0.627
## f5 ~~
## f6 0.300 0.008 39.674 0.000 0.554 0.554
##
## Intercepts:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .byconf1 0.000 0.000 0.000
## .byconf2 0.000 0.000 0.000
## .byconf3 0.000 0.000 0.000
## .byconf4 0.000 0.000 0.000
## .byloc1 0.000 0.000 0.000
## .byloc2 0.000 0.000 0.000
## .byloc3 0.000 0.000 0.000
## .byloc4 0.000 0.000 0.000
## .f1conf1 0.000 0.000 0.000
## .f1conf2 0.000 0.000 0.000
## .f1conf3 0.000 0.000 0.000
## .f1conf4 0.000 0.000 0.000
## .f1loc1 0.000 0.000 0.000
## .f1loc2 0.000 0.000 0.000
## .f1loc3 0.000 0.000 0.000
## .f1loc4 0.000 0.000 0.000
## .f2conf1 0.000 0.000 0.000
## .f2conf2 0.000 0.000 0.000
## .f2conf3 0.000 0.000 0.000
## .f2conf4 0.000 0.000 0.000
## .f2loc1 0.000 0.000 0.000
## .f2loc2 0.000 0.000 0.000
## .f2loc3 0.000 0.000 0.000
## .f2loc4 0.000 0.000 0.000
## f1 0.000 0.000 0.000
## f2 0.000 0.000 0.000
## f3 0.000 0.000 0.000
## f4 0.000 0.000 0.000
## f5 0.000 0.000 0.000
## f6 0.000 0.000 0.000
##
## Thresholds:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## byconf1|t1 -2.438 0.047 -51.980 0.000 -2.438 -2.438
## byconf1|t2 -1.484 0.021 -69.429 0.000 -1.484 -1.484
## byconf1|t3 0.399 0.014 27.635 0.000 0.399 0.399
## byconf2|t1 -2.307 0.041 -56.283 0.000 -2.307 -2.307
## byconf2|t2 -1.509 0.022 -69.550 0.000 -1.509 -1.509
## byconf2|t3 0.219 0.014 15.501 0.000 0.219 0.219
## byconf3|t1 -2.463 0.048 -51.116 0.000 -2.463 -2.463
## byconf3|t2 -1.485 0.021 -69.434 0.000 -1.485 -1.485
## byconf3|t3 0.263 0.014 18.536 0.000 0.263 0.263
## byconf4|t1 -2.203 0.037 -59.445 0.000 -2.203 -2.203
## byconf4|t2 -1.261 0.019 -66.583 0.000 -1.261 -1.261
## byconf4|t3 0.380 0.014 26.392 0.000 0.380 0.380
## byloc1|t1 -1.725 0.025 -69.015 0.000 -1.725 -1.725
## byloc1|t2 -0.948 0.017 -57.170 0.000 -0.948 -0.948
## byloc1|t3 0.416 0.014 28.743 0.000 0.416 0.416
## byloc2|t1 -2.050 0.032 -63.538 0.000 -2.050 -2.050
## byloc2|t2 -1.386 0.020 -68.566 0.000 -1.386 -1.386
## byloc2|t3 0.142 0.014 10.115 0.000 0.142 0.142
## byloc3|t1 -1.656 0.024 -69.487 0.000 -1.656 -1.656
## byloc3|t2 -0.724 0.015 -46.859 0.000 -0.724 -0.724
## byloc3|t3 0.939 0.017 56.817 0.000 0.939 0.939
## byloc4|t1 -1.742 0.025 -68.859 0.000 -1.742 -1.742
## byloc4|t2 -0.998 0.017 -59.079 0.000 -0.998 -0.998
## byloc4|t3 0.521 0.015 35.361 0.000 0.521 0.521
## f1conf1|t1 -2.285 0.040 -56.973 0.000 -2.285 -2.285
## f1conf1|t2 -1.423 0.021 -68.963 0.000 -1.423 -1.423
## f1conf1|t3 0.450 0.015 30.933 0.000 0.450 0.450
## f1conf2|t1 -2.307 0.041 -56.283 0.000 -2.307 -2.307
## f1conf2|t2 -1.474 0.021 -69.367 0.000 -1.474 -1.474
## f1conf2|t3 0.406 0.014 28.056 0.000 0.406 0.406
## f1conf3|t1 -2.432 0.047 -52.187 0.000 -2.432 -2.432
## f1conf3|t2 -1.493 0.021 -69.474 0.000 -1.493 -1.493
## f1conf3|t3 0.476 0.015 32.522 0.000 0.476 0.476
## f1conf4|t1 -2.125 0.034 -61.635 0.000 -2.125 -2.125
## f1conf4|t2 -1.105 0.018 -62.673 0.000 -1.105 -1.105
## f1conf4|t3 0.581 0.015 38.927 0.000 0.581 0.581
## f1loc1|t1 -1.816 0.027 -68.006 0.000 -1.816 -1.816
## f1loc1|t2 -0.824 0.016 -51.809 0.000 -0.824 -0.824
## f1loc1|t3 0.652 0.015 43.005 0.000 0.652 0.652
## f1loc2|t1 -2.137 0.035 -61.307 0.000 -2.137 -2.137
## f1loc2|t2 -1.315 0.019 -67.568 0.000 -1.315 -1.315
## f1loc2|t3 0.413 0.014 28.543 0.000 0.413 0.413
## f1loc3|t1 -1.904 0.029 -66.613 0.000 -1.904 -1.904
## f1loc3|t2 -0.745 0.016 -47.919 0.000 -0.745 -0.745
## f1loc3|t3 1.020 0.017 59.889 0.000 1.020 1.020
## f1loc4|t1 -1.932 0.029 -66.101 0.000 -1.932 -1.932
## f1loc4|t2 -0.910 0.016 -55.629 0.000 -0.910 -0.910
## f1loc4|t3 0.804 0.016 50.856 0.000 0.804 0.804
## f2conf1|t1 -2.330 0.042 -55.545 0.000 -2.330 -2.330
## f2conf1|t2 -1.495 0.022 -69.489 0.000 -1.495 -1.495
## f2conf1|t3 0.266 0.014 18.737 0.000 0.266 0.266
## f2conf2|t1 -2.229 0.038 -58.689 0.000 -2.229 -2.229
## f2conf2|t2 -1.545 0.022 -69.656 0.000 -1.545 -1.545
## f2conf2|t3 0.199 0.014 14.116 0.000 0.199 0.199
## f2conf3|t1 -2.381 0.044 -53.901 0.000 -2.381 -2.381
## f2conf3|t2 -1.591 0.023 -69.675 0.000 -1.591 -1.591
## f2conf3|t3 0.268 0.014 18.871 0.000 0.268 0.268
## f2conf4|t1 -2.169 0.036 -60.426 0.000 -2.169 -2.169
## f2conf4|t2 -1.244 0.019 -66.230 0.000 -1.244 -1.244
## f2conf4|t3 0.393 0.014 27.235 0.000 0.393 0.393
## f2loc1|t1 -1.700 0.025 -69.223 0.000 -1.700 -1.700
## f2loc1|t2 -0.862 0.016 -53.532 0.000 -0.862 -0.862
## f2loc1|t3 0.604 0.015 40.277 0.000 0.604 0.604
## f2loc2|t1 -2.001 0.031 -64.667 0.000 -2.001 -2.001
## f2loc2|t2 -1.346 0.020 -68.050 0.000 -1.346 -1.346
## f2loc2|t3 0.385 0.014 26.725 0.000 0.385 0.385
## f2loc3|t1 -1.829 0.027 -67.823 0.000 -1.829 -1.829
## f2loc3|t2 -0.824 0.016 -51.829 0.000 -0.824 -0.824
## f2loc3|t3 0.953 0.017 57.365 0.000 0.953 0.953
## f2loc4|t1 -1.839 0.027 -67.676 0.000 -1.839 -1.839
## f2loc4|t2 -0.954 0.017 -57.424 0.000 -0.954 -0.954
## f2loc4|t3 0.728 0.015 47.051 0.000 0.728 0.728
##
## Variances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .byconf1 0.354 0.354 0.354
## .byconf2 0.525 0.525 0.525
## .byconf3 0.606 0.606 0.606
## .byconf4 0.335 0.335 0.335
## .byloc1 0.630 0.630 0.630
## .byloc2 0.802 0.802 0.802
## .byloc3 0.562 0.562 0.562
## .byloc4 0.475 0.475 0.475
## .f1conf1 0.335 0.335 0.335
## .f1conf2 0.399 0.399 0.399
## .f1conf3 0.479 0.479 0.479
## .f1conf4 0.344 0.344 0.344
## .f1loc1 0.574 0.574 0.574
## .f1loc2 0.760 0.760 0.760
## .f1loc3 0.473 0.473 0.473
## .f1loc4 0.392 0.392 0.392
## .f2conf1 0.308 0.308 0.308
## .f2conf2 0.302 0.302 0.302
## .f2conf3 0.354 0.354 0.354
## .f2conf4 0.286 0.286 0.286
## .f2loc1 0.576 0.576 0.576
## .f2loc2 0.712 0.712 0.712
## .f2loc3 0.403 0.403 0.403
## .f2loc4 0.341 0.341 0.341
## f1 0.646 0.012 52.686 0.000 1.000 1.000
## f2 0.370 0.014 26.989 0.000 1.000 1.000
## f3 0.665 0.010 64.339 0.000 1.000 1.000
## f4 0.426 0.012 35.854 0.000 1.000 1.000
## f5 0.692 0.009 76.827 0.000 1.000 1.000
## f6 0.424 0.012 35.237 0.000 1.000 1.000
##
## Scales y*:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## byconf1 1.000 1.000 1.000
## byconf2 1.000 1.000 1.000
## byconf3 1.000 1.000 1.000
## byconf4 1.000 1.000 1.000
## byloc1 1.000 1.000 1.000
## byloc2 1.000 1.000 1.000
## byloc3 1.000 1.000 1.000
## byloc4 1.000 1.000 1.000
## f1conf1 1.000 1.000 1.000
## f1conf2 1.000 1.000 1.000
## f1conf3 1.000 1.000 1.000
## f1conf4 1.000 1.000 1.000
## f1loc1 1.000 1.000 1.000
## f1loc2 1.000 1.000 1.000
## f1loc3 1.000 1.000 1.000
## f1loc4 1.000 1.000 1.000
## f2conf1 1.000 1.000 1.000
## f2conf2 1.000 1.000 1.000
## f2conf3 1.000 1.000 1.000
## f2conf4 1.000 1.000 1.000
## f2loc1 1.000 1.000 1.000
## f2loc2 1.000 1.000 1.000
## f2loc3 1.000 1.000 1.000
## f2loc4 1.000 1.000 1.000
##
## R-Square:
## Estimate
## byconf1 0.646
## byconf2 0.475
## byconf3 0.394
## byconf4 0.665
## byloc1 0.370
## byloc2 0.198
## byloc3 0.438
## byloc4 0.525
## f1conf1 0.665
## f1conf2 0.601
## f1conf3 0.521
## f1conf4 0.656
## f1loc1 0.426
## f1loc2 0.240
## f1loc3 0.527
## f1loc4 0.608
## f2conf1 0.692
## f2conf2 0.698
## f2conf3 0.646
## f2conf4 0.714
## f2loc1 0.424
## f2loc2 0.288
## f2loc3 0.597
## f2loc4 0.6596.2 Syntax - Mplus
6.2.1 Analysis with data from single groups
TITLE: Male group
DATA: FILE IS "data\NELSMALE.txt";
TYPE IS CORRELATION STDEVIATIONS;
NOBSERVATIONS ARE 5312;
VARIABLE: NAMES ARE byconf1-byconf4 byloc1-byloc4;
ANALYSIS: ESTIMATOR=ML;
OUTPUT: SAMPSTAT STDYX RESIDUAL;
MODEL:
f1 BY byconf1-byconf4;
f2 BY byloc1-byloc4;
byconf2 WITH byconf3;## Mplus VERSION 8.4
## MUTHEN & MUTHEN
## 06/10/2021 12:19 PM
##
## INPUT INSTRUCTIONS
##
## TITLE: Male group
## DATA: FILE IS "data\NELSMALE.txt";
## TYPE IS CORRELATION STDEVIATIONS;
## NOBSERVATIONS ARE 5312;
## VARIABLE: NAMES ARE byconf1-byconf4 byloc1-byloc4;
## ANALYSIS: ESTIMATOR=ML;
## OUTPUT: SAMPSTAT STDYX RESIDUAL;
## MODEL:
## f1 BY byconf1-byconf4;
## f2 BY byloc1-byloc4;
## byconf2 WITH byconf3;
##
##
##
## 1 ERROR(S) FOUND IN THE INPUT INSTRUCTIONS
##
##
##
## Male group
##
## SUMMARY OF ANALYSIS
##
## Number of groups 1
## Number of observations 5312
##
## Number of dependent variables 8
## Number of independent variables 0
## Number of continuous latent variables 2
##
## Observed dependent variables
##
## Continuous
## BYCONF1 BYCONF2 BYCONF3 BYCONF4 BYLOC1 BYLOC2
## BYLOC3 BYLOC4
##
## Continuous latent variables
## F1 F2
##
##
## Estimator ML
## Information matrix EXPECTED
## Maximum number of iterations 1000
## Convergence criterion 0.500D-04
## Maximum number of steepest descent iterations 20
##
## Input data file(s)
## data\NELSMALE.txt
##
## Input data format FREE
##
##
## SAMPLE STATISTICS
##
##
## SAMPLE STATISTICS
##
##
## Covariances/Correlations/Residual Correlations
## BYCONF1 BYCONF2 BYCONF3 BYCONF4 BYLOC1
## ________ ________ ________ ________ ________
## BYCONF1 0.343
## BYCONF2 0.125 0.413
## BYCONF3 0.101 0.166 0.384
## BYCONF4 0.184 0.155 0.126 0.426
## BYLOC1 0.096 0.084 0.070 0.119 0.646
## BYLOC2 0.025 0.077 0.045 0.060 0.146
## BYLOC3 0.093 0.071 0.065 0.118 0.185
## BYLOC4 0.095 0.093 0.082 0.130 0.180
##
##
## Covariances/Correlations/Residual Correlations
## BYLOC2 BYLOC3 BYLOC4
## ________ ________ ________
## BYLOC2 0.560
## BYLOC3 0.112 0.582
## BYLOC4 0.135 0.234 0.587
##
##
## THE MODEL ESTIMATION TERMINATED NORMALLY
##
##
##
## MODEL FIT INFORMATION
##
## Number of Free Parameters 18
##
## Loglikelihood
##
## H0 Value -41500.792
## H1 Value -41427.307
##
## Information Criteria
##
## Akaike (AIC) 83037.584
## Bayesian (BIC) 83155.983
## Sample-Size Adjusted BIC 83098.785
## (n* = (n + 2) / 24)
##
## Chi-Square Test of Model Fit
##
## Value 146.970
## Degrees of Freedom 18
## P-Value 0.0000
##
## RMSEA (Root Mean Square Error Of Approximation)
##
## Estimate 0.037
## 90 Percent C.I. 0.031 0.042
## Probability RMSEA <= .05 1.000
##
## CFI/TLI
##
## CFI 0.981
## TLI 0.970
##
## Chi-Square Test of Model Fit for the Baseline Model
##
## Value 6651.699
## Degrees of Freedom 28
## P-Value 0.0000
##
## SRMR (Standardized Root Mean Square Residual)
##
## Value 0.022
##
##
##
## MODEL RESULTS
##
## Two-Tailed
## Estimate S.E. Est./S.E. P-Value
##
## F1 BY
## BYCONF1 1.000 0.000 999.000 999.000
## BYCONF2 0.863 0.031 27.486 0.000
## BYCONF3 0.701 0.029 23.820 0.000
## BYCONF4 1.272 0.040 31.424 0.000
##
## F2 BY
## BYLOC1 1.000 0.000 999.000 999.000
## BYLOC2 0.667 0.036 18.716 0.000
## BYLOC3 1.115 0.045 24.711 0.000
## BYLOC4 1.179 0.047 24.971 0.000
##
## F2 WITH
## F1 0.085 0.004 18.978 0.000
##
## BYCONF2 WITH
## BYCONF3 0.079 0.005 15.304 0.000
##
## Variances
## F1 0.144 0.007 21.119 0.000
## F2 0.169 0.011 15.846 0.000
##
## Residual Variances
## BYCONF1 0.200 0.006 35.342 0.000
## BYCONF2 0.306 0.007 43.856 0.000
## BYCONF3 0.314 0.007 46.432 0.000
## BYCONF4 0.194 0.007 25.998 0.000
## BYLOC1 0.477 0.011 42.099 0.000
## BYLOC2 0.484 0.010 47.493 0.000
## BYLOC3 0.371 0.010 36.188 0.000
## BYLOC4 0.351 0.010 33.481 0.000
##
##
## STANDARDIZED MODEL RESULTS
##
##
## STDYX Standardization
##
## Two-Tailed
## Estimate S.E. Est./S.E. P-Value
##
## F1 BY
## BYCONF1 0.647 0.012 52.931 0.000
## BYCONF2 0.509 0.013 38.003 0.000
## BYCONF3 0.428 0.014 29.922 0.000
## BYCONF4 0.738 0.012 61.561 0.000
##
## F2 BY
## BYLOC1 0.512 0.014 36.211 0.000
## BYLOC2 0.367 0.015 23.994 0.000
## BYLOC3 0.602 0.014 44.159 0.000
## BYLOC4 0.634 0.014 46.896 0.000
##
## F2 WITH
## F1 0.546 0.017 31.929 0.000
##
## BYCONF2 WITH
## BYCONF3 0.255 0.014 17.679 0.000
##
## Variances
## F1 1.000 0.000 999.000 999.000
## F2 1.000 0.000 999.000 999.000
##
## Residual Variances
## BYCONF1 0.582 0.016 36.780 0.000
## BYCONF2 0.741 0.014 54.354 0.000
## BYCONF3 0.817 0.012 66.598 0.000
## BYCONF4 0.455 0.018 25.706 0.000
## BYLOC1 0.738 0.014 50.956 0.000
## BYLOC2 0.865 0.011 77.096 0.000
## BYLOC3 0.638 0.016 38.937 0.000
## BYLOC4 0.599 0.017 34.968 0.000
##
##
## R-SQUARE
##
## Observed Two-Tailed
## Variable Estimate S.E. Est./S.E. P-Value
##
## BYCONF1 0.418 0.016 26.465 0.000
## BYCONF2 0.259 0.014 19.002 0.000
## BYCONF3 0.183 0.012 14.961 0.000
## BYCONF4 0.545 0.018 30.781 0.000
## BYLOC1 0.262 0.014 18.106 0.000
## BYLOC2 0.135 0.011 11.997 0.000
## BYLOC3 0.362 0.016 22.080 0.000
## BYLOC4 0.401 0.017 23.448 0.000
##
##
## QUALITY OF NUMERICAL RESULTS
##
## Condition Number for the Information Matrix 0.134E-02
## (ratio of smallest to largest eigenvalue)
##
##
## RESIDUAL OUTPUT
##
##
## ESTIMATED MODEL AND RESIDUALS (OBSERVED - ESTIMATED)
##
##
## Model Estimated Covariances/Correlations/Residual Correlations
## BYCONF1 BYCONF2 BYCONF3 BYCONF4 BYLOC1
## ________ ________ ________ ________ ________
## BYCONF1 0.343
## BYCONF2 0.124 0.413
## BYCONF3 0.101 0.166 0.384
## BYCONF4 0.183 0.158 0.128 0.426
## BYLOC1 0.085 0.074 0.060 0.108 0.646
## BYLOC2 0.057 0.049 0.040 0.072 0.113
## BYLOC3 0.095 0.082 0.067 0.121 0.189
## BYLOC4 0.100 0.087 0.070 0.128 0.200
##
##
## Model Estimated Covariances/Correlations/Residual Correlations
## BYLOC2 BYLOC3 BYLOC4
## ________ ________ ________
## BYLOC2 0.559
## BYLOC3 0.126 0.582
## BYLOC4 0.133 0.223 0.587
##
##
## Residuals for Covariances/Correlations/Residual Correlations
## BYCONF1 BYCONF2 BYCONF3 BYCONF4 BYLOC1
## ________ ________ ________ ________ ________
## BYCONF1 0.000
## BYCONF2 0.001 0.000
## BYCONF3 0.000 0.000 0.000
## BYCONF4 0.002 -0.002 -0.002 0.000
## BYLOC1 0.010 0.010 0.011 0.011 0.000
## BYLOC2 -0.032 0.028 0.005 -0.012 0.033
## BYLOC3 -0.002 -0.011 -0.002 -0.003 -0.004
## BYLOC4 -0.006 0.006 0.011 0.002 -0.019
##
##
## Residuals for Covariances/Correlations/Residual Correlations
## BYLOC2 BYLOC3 BYLOC4
## ________ ________ ________
## BYLOC2 0.000
## BYLOC3 -0.014 0.000
## BYLOC4 0.001 0.012 0.000
##
##
## Standardized Residuals (z-scores) for Covariances/Correlations/Residual Corr
## BYCONF1 BYCONF2 BYCONF3 BYCONF4 BYLOC1
## ________ ________ ________ ________ ________
## BYCONF1 0.000
## BYCONF2 0.322 0.000
## BYCONF3 -0.008 0.000 0.000
## BYCONF4 2.016 -1.557 -1.363 0.000
## BYLOC1 2.166 1.737 1.797 2.201 0.000
## BYLOC2 -6.483 4.824 0.818 -2.340 5.737
## BYLOC3 -0.586 -2.177 -0.321 -0.788 -1.031
## BYLOC4 -1.443 1.147 2.148 0.617 -6.355
##
##
## Standardized Residuals (z-scores) for Covariances/Correlations/Residual Corr
## BYLOC2 BYLOC3 BYLOC4
## ________ ________ ________
## BYLOC2 0.000
## BYLOC3 -3.261 0.086
## BYLOC4 0.353 4.623 0.087
##
##
## Normalized Residuals for Covariances/Correlations/Residual Correlations
## BYCONF1 BYCONF2 BYCONF3 BYCONF4 BYLOC1
## ________ ________ ________ ________ ________
## BYCONF1 0.000
## BYCONF2 0.122 0.000
## BYCONF3 -0.004 0.000 0.000
## BYCONF4 0.294 -0.390 -0.429 0.000
## BYLOC1 1.587 1.422 1.538 1.472 0.000
## BYLOC2 -5.280 4.250 0.741 -1.797 3.833
## BYLOC3 -0.384 -1.667 -0.261 -0.451 -0.419
## BYLOC4 -0.905 0.860 1.717 0.334 -2.196
##
##
## Normalized Residuals for Covariances/Correlations/Residual Correlations
## BYLOC2 BYLOC3 BYLOC4
## ________ ________ ________
## BYLOC2 0.000
## BYLOC3 -1.770 0.000
## BYLOC4 0.178 1.345 0.000
##
##
## Beginning Time: 12:19:50
## Ending Time: 12:19:50
## Elapsed Time: 00:00:00
##
##
##
## MUTHEN & MUTHEN
## 3463 Stoner Ave.
## Los Angeles, CA 90066
##
## Tel: (310) 391-9971
## Fax: (310) 391-8971
## Web: www.StatModel.com
## Support: Support@StatModel.com
##
## Copyright (c) 1998-2019 Muthen & MuthenTITLE: Female group
DATA: FILE IS "data\NELSFEMALE.txt";
TYPE IS CORRELATION STDEVIATIONS;
NOBSERVATIONS ARE 6008;
VARIABLE: NAMES ARE byconf1-byconf4 byloc1-byloc4;
ANALYSIS: ESTIMATOR=ML;
OUTPUT: SAMPSTAT STDYX RESIDUAL;
MODEL:
f1 BY byconf1-byconf4;
f2 BY byloc1-byloc4;
byconf2 WITH byconf3;## Mplus VERSION 8.4
## MUTHEN & MUTHEN
## 06/10/2021 12:19 PM
##
## INPUT INSTRUCTIONS
##
## TITLE: Female group
## DATA: FILE IS "data\NELSFEMALE.txt";
## TYPE IS CORRELATION STDEVIATIONS;
## NOBSERVATIONS ARE 6008;
## VARIABLE: NAMES ARE byconf1-byconf4 byloc1-byloc4;
## ANALYSIS: ESTIMATOR=ML;
## OUTPUT: SAMPSTAT STDYX RESIDUAL;
## MODEL:
## f1 BY byconf1-byconf4;
## f2 BY byloc1-byloc4;
## byconf2 WITH byconf3;
##
##
##
## 1 ERROR(S) FOUND IN THE INPUT INSTRUCTIONS
##
##
##
## Female group
##
## SUMMARY OF ANALYSIS
##
## Number of groups 1
## Number of observations 6008
##
## Number of dependent variables 8
## Number of independent variables 0
## Number of continuous latent variables 2
##
## Observed dependent variables
##
## Continuous
## BYCONF1 BYCONF2 BYCONF3 BYCONF4 BYLOC1 BYLOC2
## BYLOC3 BYLOC4
##
## Continuous latent variables
## F1 F2
##
##
## Estimator ML
## Information matrix EXPECTED
## Maximum number of iterations 1000
## Convergence criterion 0.500D-04
## Maximum number of steepest descent iterations 20
##
## Input data file(s)
## data\NELSFEMALE.txt
##
## Input data format FREE
##
##
## SAMPLE STATISTICS
##
##
## SAMPLE STATISTICS
##
##
## Covariances/Correlations/Residual Correlations
## BYCONF1 BYCONF2 BYCONF3 BYCONF4 BYLOC1
## ________ ________ ________ ________ ________
## BYCONF1 0.379
## BYCONF2 0.167 0.423
## BYCONF3 0.134 0.183 0.412
## BYCONF4 0.248 0.194 0.162 0.480
## BYLOC1 0.111 0.109 0.087 0.157 0.648
## BYLOC2 0.032 0.072 0.055 0.076 0.156
## BYLOC3 0.108 0.097 0.093 0.147 0.205
## BYLOC4 0.132 0.129 0.110 0.184 0.229
##
##
## Covariances/Correlations/Residual Correlations
## BYLOC2 BYLOC3 BYLOC4
## ________ ________ ________
## BYLOC2 0.494
## BYLOC3 0.122 0.569
## BYLOC4 0.140 0.279 0.634
##
##
## THE MODEL ESTIMATION TERMINATED NORMALLY
##
##
##
## MODEL FIT INFORMATION
##
## Number of Free Parameters 18
##
## Loglikelihood
##
## H0 Value -46343.263
## H1 Value -46244.110
##
## Information Criteria
##
## Akaike (AIC) 92722.527
## Bayesian (BIC) 92843.142
## Sample-Size Adjusted BIC 92785.943
## (n* = (n + 2) / 24)
##
## Chi-Square Test of Model Fit
##
## Value 198.307
## Degrees of Freedom 18
## P-Value 0.0000
##
## RMSEA (Root Mean Square Error Of Approximation)
##
## Estimate 0.041
## 90 Percent C.I. 0.036 0.046
## Probability RMSEA <= .05 0.998
##
## CFI/TLI
##
## CFI 0.982
## TLI 0.972
##
## Chi-Square Test of Model Fit for the Baseline Model
##
## Value 10198.456
## Degrees of Freedom 28
## P-Value 0.0000
##
## SRMR (Standardized Root Mean Square Residual)
##
## Value 0.024
##
##
##
## MODEL RESULTS
##
## Two-Tailed
## Estimate S.E. Est./S.E. P-Value
##
## F1 BY
## BYCONF1 1.000 0.000 999.000 999.000
## BYCONF2 0.820 0.022 36.510 0.000
## BYCONF3 0.677 0.022 30.855 0.000
## BYCONF4 1.240 0.028 43.702 0.000
##
## F2 BY
## BYLOC1 1.000 0.000 999.000 999.000
## BYLOC2 0.604 0.027 22.121 0.000
## BYLOC3 1.091 0.035 30.827 0.000
## BYLOC4 1.261 0.040 31.500 0.000
##
## F2 WITH
## F1 0.112 0.005 23.313 0.000
##
## BYCONF2 WITH
## BYCONF3 0.073 0.005 15.686 0.000
##
## Variances
## F1 0.198 0.007 27.642 0.000
## F2 0.195 0.010 19.176 0.000
##
## Residual Variances
## BYCONF1 0.181 0.005 36.090 0.000
## BYCONF2 0.289 0.006 47.311 0.000
## BYCONF3 0.321 0.006 50.052 0.000
## BYCONF4 0.176 0.007 26.935 0.000
## BYLOC1 0.453 0.010 45.301 0.000
## BYLOC2 0.423 0.008 51.184 0.000
## BYLOC3 0.336 0.009 39.152 0.000
## BYLOC4 0.324 0.010 33.421 0.000
##
##
## STANDARDIZED MODEL RESULTS
##
##
## STDYX Standardization
##
## Two-Tailed
## Estimate S.E. Est./S.E. P-Value
##
## F1 BY
## BYCONF1 0.722 0.009 78.003 0.000
## BYCONF2 0.561 0.011 51.135 0.000
## BYCONF3 0.469 0.012 38.725 0.000
## BYCONF4 0.796 0.009 90.365 0.000
##
## F2 BY
## BYLOC1 0.548 0.012 45.633 0.000
## BYLOC2 0.379 0.014 27.953 0.000
## BYLOC3 0.639 0.011 56.697 0.000
## BYLOC4 0.699 0.011 64.081 0.000
##
## F2 WITH
## F1 0.571 0.014 41.129 0.000
##
## BYCONF2 WITH
## BYCONF3 0.240 0.013 17.812 0.000
##
## Variances
## F1 1.000 0.000 999.000 999.000
## F2 1.000 0.000 999.000 999.000
##
## Residual Variances
## BYCONF1 0.478 0.013 35.755 0.000
## BYCONF2 0.685 0.012 55.589 0.000
## BYCONF3 0.780 0.011 68.526 0.000
## BYCONF4 0.366 0.014 26.104 0.000
## BYLOC1 0.700 0.013 53.115 0.000
## BYLOC2 0.856 0.010 83.155 0.000
## BYLOC3 0.592 0.014 41.130 0.000
## BYLOC4 0.512 0.015 33.566 0.000
##
##
## R-SQUARE
##
## Observed Two-Tailed
## Variable Estimate S.E. Est./S.E. P-Value
##
## BYCONF1 0.522 0.013 39.002 0.000
## BYCONF2 0.315 0.012 25.568 0.000
## BYCONF3 0.220 0.011 19.362 0.000
## BYCONF4 0.634 0.014 45.182 0.000
## BYLOC1 0.300 0.013 22.816 0.000
## BYLOC2 0.144 0.010 13.976 0.000
## BYLOC3 0.408 0.014 28.349 0.000
## BYLOC4 0.488 0.015 32.040 0.000
##
##
## QUALITY OF NUMERICAL RESULTS
##
## Condition Number for the Information Matrix 0.201E-02
## (ratio of smallest to largest eigenvalue)
##
##
## RESIDUAL OUTPUT
##
##
## ESTIMATED MODEL AND RESIDUALS (OBSERVED - ESTIMATED)
##
##
## Model Estimated Covariances/Correlations/Residual Correlations
## BYCONF1 BYCONF2 BYCONF3 BYCONF4 BYLOC1
## ________ ________ ________ ________ ________
## BYCONF1 0.379
## BYCONF2 0.162 0.422
## BYCONF3 0.134 0.183 0.412
## BYCONF4 0.245 0.201 0.166 0.480
## BYLOC1 0.112 0.092 0.076 0.139 0.648
## BYLOC2 0.068 0.056 0.046 0.084 0.118
## BYLOC3 0.122 0.100 0.083 0.152 0.212
## BYLOC4 0.141 0.116 0.096 0.175 0.245
##
##
## Model Estimated Covariances/Correlations/Residual Correlations
## BYLOC2 BYLOC3 BYLOC4
## ________ ________ ________
## BYLOC2 0.494
## BYLOC3 0.128 0.568
## BYLOC4 0.148 0.268 0.634
##
##
## Residuals for Covariances/Correlations/Residual Correlations
## BYCONF1 BYCONF2 BYCONF3 BYCONF4 BYLOC1
## ________ ________ ________ ________ ________
## BYCONF1 0.000
## BYCONF2 0.005 0.000
## BYCONF3 0.000 0.000 0.000
## BYCONF4 0.003 -0.007 -0.004 0.000
## BYLOC1 -0.002 0.017 0.011 0.018 0.000
## BYLOC2 -0.035 0.017 0.009 -0.009 0.039
## BYLOC3 -0.014 -0.004 0.010 -0.005 -0.007
## BYLOC4 -0.009 0.013 0.014 0.008 -0.016
##
##
## Residuals for Covariances/Correlations/Residual Correlations
## BYLOC2 BYLOC3 BYLOC4
## ________ ________ ________
## BYLOC2 0.000
## BYLOC3 -0.006 0.000
## BYLOC4 -0.008 0.011 0.000
##
##
## Standardized Residuals (z-scores) for Covariances/Correlations/Residual Corr
## BYCONF1 BYCONF2 BYCONF3 BYCONF4 BYLOC1
## ________ ________ ________ ________ ________
## BYCONF1 999.000
## BYCONF2 2.764 0.062
## BYCONF3 0.182 0.110 0.111
## BYCONF4 3.412 -6.033 -2.642 0.120
## BYLOC1 -0.351 3.186 2.016 3.869 0.074
## BYLOC2 -8.072 3.225 1.658 -1.805 7.682
## BYLOC3 -3.892 -0.806 1.995 -1.344 -2.342
## BYLOC4 -2.689 2.670 2.750 2.381 -6.424
##
##
## Standardized Residuals (z-scores) for Covariances/Correlations/Residual Corr
## BYLOC2 BYLOC3 BYLOC4
## ________ ________ ________
## BYLOC2 0.059
## BYLOC3 -1.594 0.000
## BYLOC4 -2.384 5.257 0.000
##
##
## Normalized Residuals for Covariances/Correlations/Residual Correlations
## BYCONF1 BYCONF2 BYCONF3 BYCONF4 BYLOC1
## ________ ________ ________ ________ ________
## BYCONF1 0.000
## BYCONF2 0.900 0.000
## BYCONF3 0.072 0.000 0.000
## BYCONF4 0.400 -1.128 -0.704 0.001
## BYLOC1 -0.239 2.523 1.684 2.442 0.000
## BYLOC2 -6.299 2.786 1.485 -1.340 5.086
## BYLOC3 -2.305 -0.592 1.579 -0.707 -0.892
## BYLOC4 -1.446 1.877 2.095 1.119 -1.830
##
##
## Normalized Residuals for Covariances/Correlations/Residual Correlations
## BYLOC2 BYLOC3 BYLOC4
## ________ ________ ________
## BYLOC2 0.000
## BYLOC3 -0.854 0.000
## BYLOC4 -1.059 1.308 0.000
##
##
## Beginning Time: 12:19:50
## Ending Time: 12:19:51
## Elapsed Time: 00:00:01
##
##
##
## MUTHEN & MUTHEN
## 3463 Stoner Ave.
## Los Angeles, CA 90066
##
## Tel: (310) 391-9971
## Fax: (310) 391-8971
## Web: www.StatModel.com
## Support: Support@StatModel.com
##
## Copyright (c) 1998-2019 Muthen & Muthen6.2.2 Analysis with data from both groups
6.2.2.1 both groups, no constraints
TITLE: both groups, no constraints
DATA: FILE IS "data\NELSTWOGROUPS.txt";
TYPE IS CORRELATION STDEVIATIONS;
NOBSERVATIONS ARE 5312 6008;
NGROUPS IS 2;
VARIABLE: NAMES ARE byconf1-byconf4 byloc1-byloc4;
ANALYSIS: ESTIMATOR=ML;
OUTPUT: SAMPSTAT STDYX RESIDUAL;
MODEL:
f1 BY byconf1-byconf4;
f2 BY byloc1-byloc4;
f1 WITH f2;
byconf2 WITH byconf3;
MODEL g2:
f1 BY byconf2-byconf4;
f2 BY byloc2-byloc4;
f1 WITH f2;
byconf2 WITH byconf3;## Mplus VERSION 8.4
## MUTHEN & MUTHEN
## 06/10/2021 12:19 PM
##
## INPUT INSTRUCTIONS
##
## TITLE: both groups, no constraints
## DATA: FILE IS "data\NELSTWOGROUPS.txt";
## TYPE IS CORRELATION STDEVIATIONS;
## NOBSERVATIONS ARE 5312 6008;
## NGROUPS IS 2;
## VARIABLE: NAMES ARE byconf1-byconf4 byloc1-byloc4;
## ANALYSIS: ESTIMATOR=ML;
## OUTPUT: SAMPSTAT STDYX RESIDUAL;
## MODEL:
## f1 BY byconf1-byconf4;
## f2 BY byloc1-byloc4;
## f1 WITH f2;
## byconf2 WITH byconf3;
## MODEL g2:
## f1 BY byconf2-byconf4;
## f2 BY byloc2-byloc4;
## f1 WITH f2;
## byconf2 WITH byconf3;
##
##
##
## 1 ERROR(S) FOUND IN THE INPUT INSTRUCTIONS
##
##
##
## both groups, no constraints
##
## SUMMARY OF ANALYSIS
##
## Number of groups 2
## Number of observations
## Group G1 5312
## Group G2 6008
## Total sample size 11320
##
## Number of dependent variables 8
## Number of independent variables 0
## Number of continuous latent variables 2
##
## Observed dependent variables
##
## Continuous
## BYCONF1 BYCONF2 BYCONF3 BYCONF4 BYLOC1 BYLOC2
## BYLOC3 BYLOC4
##
## Continuous latent variables
## F1 F2
##
##
## Estimator ML
## Information matrix EXPECTED
## Maximum number of iterations 1000
## Convergence criterion 0.500D-04
## Maximum number of steepest descent iterations 20
##
## Input data file(s)
## data\NELSTWOGROUPS.txt
##
## Input data format FREE
##
##
## SAMPLE STATISTICS
##
##
## SAMPLE STATISTICS FOR G1
##
##
## Covariances/Correlations/Residual Correlations
## BYCONF1 BYCONF2 BYCONF3 BYCONF4 BYLOC1
## ________ ________ ________ ________ ________
## BYCONF1 0.343
## BYCONF2 0.125 0.413
## BYCONF3 0.101 0.166 0.384
## BYCONF4 0.184 0.155 0.126 0.426
## BYLOC1 0.096 0.084 0.070 0.119 0.646
## BYLOC2 0.025 0.077 0.045 0.060 0.146
## BYLOC3 0.093 0.071 0.065 0.118 0.185
## BYLOC4 0.095 0.093 0.082 0.130 0.180
##
##
## Covariances/Correlations/Residual Correlations
## BYLOC2 BYLOC3 BYLOC4
## ________ ________ ________
## BYLOC2 0.560
## BYLOC3 0.112 0.582
## BYLOC4 0.135 0.234 0.587
##
##
## SAMPLE STATISTICS FOR G2
##
##
## Covariances/Correlations/Residual Correlations
## BYCONF1 BYCONF2 BYCONF3 BYCONF4 BYLOC1
## ________ ________ ________ ________ ________
## BYCONF1 0.379
## BYCONF2 0.167 0.423
## BYCONF3 0.134 0.183 0.412
## BYCONF4 0.248 0.194 0.162 0.480
## BYLOC1 0.111 0.109 0.087 0.157 0.648
## BYLOC2 0.032 0.072 0.055 0.076 0.156
## BYLOC3 0.108 0.097 0.093 0.147 0.205
## BYLOC4 0.132 0.129 0.110 0.184 0.229
##
##
## Covariances/Correlations/Residual Correlations
## BYLOC2 BYLOC3 BYLOC4
## ________ ________ ________
## BYLOC2 0.494
## BYLOC3 0.122 0.569
## BYLOC4 0.140 0.279 0.634
##
##
## THE MODEL ESTIMATION TERMINATED NORMALLY
##
##
##
## MODEL FIT INFORMATION
##
## Number of Free Parameters 36
##
## Loglikelihood
##
## H0 Value -87844.056
## H1 Value -87671.417
##
## Information Criteria
##
## Akaike (AIC) 175760.111
## Bayesian (BIC) 176024.147
## Sample-Size Adjusted BIC 175909.743
## (n* = (n + 2) / 24)
##
## Chi-Square Test of Model Fit
##
## Value 345.277
## Degrees of Freedom 36
## P-Value 0.0000
##
## Chi-Square Contribution From Each Group
##
## G1 146.970
## G2 198.307
##
## RMSEA (Root Mean Square Error Of Approximation)
##
## Estimate 0.039
## 90 Percent C.I. 0.035 0.043
## Probability RMSEA <= .05 1.000
##
## CFI/TLI
##
## CFI 0.982
## TLI 0.971
##
## Chi-Square Test of Model Fit for the Baseline Model
##
## Value 16850.155
## Degrees of Freedom 56
## P-Value 0.0000
##
## SRMR (Standardized Root Mean Square Residual)
##
## Value 0.023
##
##
##
## MODEL RESULTS
##
## Two-Tailed
## Estimate S.E. Est./S.E. P-Value
##
## Group G1
##
## F1 BY
## BYCONF1 1.000 0.000 999.000 999.000
## BYCONF2 0.863 0.031 27.486 0.000
## BYCONF3 0.701 0.029 23.820 0.000
## BYCONF4 1.272 0.040 31.424 0.000
##
## F2 BY
## BYLOC1 1.000 0.000 999.000 999.000
## BYLOC2 0.667 0.036 18.716 0.000
## BYLOC3 1.115 0.045 24.712 0.000
## BYLOC4 1.179 0.047 24.971 0.000
##
## F1 WITH
## F2 0.085 0.004 18.978 0.000
##
## BYCONF2 WITH
## BYCONF3 0.079 0.005 15.304 0.000
##
## Variances
## F1 0.144 0.007 21.119 0.000
## F2 0.169 0.011 15.846 0.000
##
## Residual Variances
## BYCONF1 0.200 0.006 35.342 0.000
## BYCONF2 0.306 0.007 43.856 0.000
## BYCONF3 0.314 0.007 46.432 0.000
## BYCONF4 0.194 0.007 25.998 0.000
## BYLOC1 0.477 0.011 42.099 0.000
## BYLOC2 0.484 0.010 47.493 0.000
## BYLOC3 0.371 0.010 36.188 0.000
## BYLOC4 0.351 0.010 33.481 0.000
##
## Group G2
##
## F1 BY
## BYCONF1 1.000 0.000 999.000 999.000
## BYCONF2 0.820 0.022 36.510 0.000
## BYCONF3 0.677 0.022 30.855 0.000
## BYCONF4 1.240 0.028 43.702 0.000
##
## F2 BY
## BYLOC1 1.000 0.000 999.000 999.000
## BYLOC2 0.604 0.027 22.120 0.000
## BYLOC3 1.091 0.035 30.827 0.000
## BYLOC4 1.261 0.040 31.500 0.000
##
## F1 WITH
## F2 0.112 0.005 23.313 0.000
##
## BYCONF2 WITH
## BYCONF3 0.073 0.005 15.686 0.000
##
## Variances
## F1 0.198 0.007 27.643 0.000
## F2 0.195 0.010 19.176 0.000
##
## Residual Variances
## BYCONF1 0.181 0.005 36.089 0.000
## BYCONF2 0.289 0.006 47.311 0.000
## BYCONF3 0.321 0.006 50.052 0.000
## BYCONF4 0.176 0.007 26.935 0.000
## BYLOC1 0.453 0.010 45.301 0.000
## BYLOC2 0.423 0.008 51.184 0.000
## BYLOC3 0.336 0.009 39.151 0.000
## BYLOC4 0.324 0.010 33.420 0.000
##
##
## STANDARDIZED MODEL RESULTS
##
##
## STDYX Standardization
##
## Two-Tailed
## Estimate S.E. Est./S.E. P-Value
##
## Group G1
##
## F1 BY
## BYCONF1 0.647 0.012 52.931 0.000
## BYCONF2 0.509 0.013 38.003 0.000
## BYCONF3 0.428 0.014 29.922 0.000
## BYCONF4 0.738 0.012 61.562 0.000
##
## F2 BY
## BYLOC1 0.512 0.014 36.212 0.000
## BYLOC2 0.367 0.015 23.994 0.000
## BYLOC3 0.602 0.014 44.159 0.000
## BYLOC4 0.634 0.014 46.895 0.000
##
## F1 WITH
## F2 0.546 0.017 31.929 0.000
##
## BYCONF2 WITH
## BYCONF3 0.255 0.014 17.679 0.000
##
## Variances
## F1 1.000 0.000 999.000 999.000
## F2 1.000 0.000 999.000 999.000
##
## Residual Variances
## BYCONF1 0.582 0.016 36.781 0.000
## BYCONF2 0.741 0.014 54.354 0.000
## BYCONF3 0.817 0.012 66.598 0.000
## BYCONF4 0.455 0.018 25.706 0.000
## BYLOC1 0.738 0.014 50.955 0.000
## BYLOC2 0.865 0.011 77.096 0.000
## BYLOC3 0.638 0.016 38.938 0.000
## BYLOC4 0.599 0.017 34.968 0.000
##
## Group G2
##
## F1 BY
## BYCONF1 0.722 0.009 78.005 0.000
## BYCONF2 0.561 0.011 51.134 0.000
## BYCONF3 0.469 0.012 38.725 0.000
## BYCONF4 0.796 0.009 90.364 0.000
##
## F2 BY
## BYLOC1 0.548 0.012 45.633 0.000
## BYLOC2 0.379 0.014 27.952 0.000
## BYLOC3 0.639 0.011 56.698 0.000
## BYLOC4 0.699 0.011 64.082 0.000
##
## F1 WITH
## F2 0.571 0.014 41.128 0.000
##
## BYCONF2 WITH
## BYCONF3 0.240 0.013 17.813 0.000
##
## Variances
## F1 1.000 0.000 999.000 999.000
## F2 1.000 0.000 999.000 999.000
##
## Residual Variances
## BYCONF1 0.478 0.013 35.754 0.000
## BYCONF2 0.685 0.012 55.589 0.000
## BYCONF3 0.780 0.011 68.526 0.000
## BYCONF4 0.366 0.014 26.105 0.000
## BYLOC1 0.700 0.013 53.116 0.000
## BYLOC2 0.856 0.010 83.157 0.000
## BYLOC3 0.592 0.014 41.129 0.000
## BYLOC4 0.512 0.015 33.565 0.000
##
##
## R-SQUARE
##
## Group G1
##
## Observed Two-Tailed
## Variable Estimate S.E. Est./S.E. P-Value
##
## BYCONF1 0.418 0.016 26.465 0.000
## BYCONF2 0.259 0.014 19.002 0.000
## BYCONF3 0.183 0.012 14.961 0.000
## BYCONF4 0.545 0.018 30.781 0.000
## BYLOC1 0.262 0.014 18.106 0.000
## BYLOC2 0.135 0.011 11.997 0.000
## BYLOC3 0.362 0.016 22.080 0.000
## BYLOC4 0.401 0.017 23.447 0.000
##
## Group G2
##
## Observed Two-Tailed
## Variable Estimate S.E. Est./S.E. P-Value
##
## BYCONF1 0.522 0.013 39.002 0.000
## BYCONF2 0.315 0.012 25.567 0.000
## BYCONF3 0.220 0.011 19.363 0.000
## BYCONF4 0.634 0.014 45.182 0.000
## BYLOC1 0.300 0.013 22.816 0.000
## BYLOC2 0.144 0.010 13.976 0.000
## BYLOC3 0.408 0.014 28.349 0.000
## BYLOC4 0.488 0.015 32.041 0.000
##
##
## QUALITY OF NUMERICAL RESULTS
##
## Condition Number for the Information Matrix 0.134E-02
## (ratio of smallest to largest eigenvalue)
##
##
## RESIDUAL OUTPUT
##
##
## ESTIMATED MODEL AND RESIDUALS (OBSERVED - ESTIMATED) FOR G1
##
##
## Model Estimated Covariances/Correlations/Residual Correlations
## BYCONF1 BYCONF2 BYCONF3 BYCONF4 BYLOC1
## ________ ________ ________ ________ ________
## BYCONF1 0.343
## BYCONF2 0.124 0.413
## BYCONF3 0.101 0.166 0.384
## BYCONF4 0.183 0.158 0.128 0.426
## BYLOC1 0.085 0.074 0.060 0.108 0.646
## BYLOC2 0.057 0.049 0.040 0.072 0.113
## BYLOC3 0.095 0.082 0.067 0.121 0.189
## BYLOC4 0.100 0.087 0.070 0.128 0.200
##
##
## Model Estimated Covariances/Correlations/Residual Correlations
## BYLOC2 BYLOC3 BYLOC4
## ________ ________ ________
## BYLOC2 0.559
## BYLOC3 0.126 0.582
## BYLOC4 0.133 0.223 0.587
##
##
## Residuals for Covariances/Correlations/Residual Correlations
## BYCONF1 BYCONF2 BYCONF3 BYCONF4 BYLOC1
## ________ ________ ________ ________ ________
## BYCONF1 0.000
## BYCONF2 0.001 0.000
## BYCONF3 0.000 0.000 0.000
## BYCONF4 0.002 -0.002 -0.002 0.000
## BYLOC1 0.010 0.010 0.011 0.011 0.000
## BYLOC2 -0.032 0.028 0.005 -0.012 0.033
## BYLOC3 -0.002 -0.011 -0.002 -0.003 -0.004
## BYLOC4 -0.006 0.006 0.011 0.002 -0.019
##
##
## Residuals for Covariances/Correlations/Residual Correlations
## BYLOC2 BYLOC3 BYLOC4
## ________ ________ ________
## BYLOC2 0.000
## BYLOC3 -0.014 0.000
## BYLOC4 0.001 0.012 0.000
##
##
## Standardized Residuals (z-scores) for Covariances/Correlations/Residual Corr
## BYCONF1 BYCONF2 BYCONF3 BYCONF4 BYLOC1
## ________ ________ ________ ________ ________
## BYCONF1 0.000
## BYCONF2 0.323 0.000
## BYCONF3 -0.008 0.000 0.000
## BYCONF4 2.016 -1.558 -1.364 0.000
## BYLOC1 2.165 1.737 1.797 2.200 0.000
## BYLOC2 -6.483 4.824 0.818 -2.340 5.736
## BYLOC3 -0.586 -2.177 -0.322 -0.789 -1.032
## BYLOC4 -1.443 1.147 2.148 0.617 -6.355
##
##
## Standardized Residuals (z-scores) for Covariances/Correlations/Residual Corr
## BYLOC2 BYLOC3 BYLOC4
## ________ ________ ________
## BYLOC2 0.051
## BYLOC3 -3.260 0.121
## BYLOC4 0.354 4.624 0.122
##
##
## Normalized Residuals for Covariances/Correlations/Residual Correlations
## BYCONF1 BYCONF2 BYCONF3 BYCONF4 BYLOC1
## ________ ________ ________ ________ ________
## BYCONF1 0.000
## BYCONF2 0.122 0.000
## BYCONF3 -0.004 0.000 0.000
## BYCONF4 0.294 -0.390 -0.429 0.000
## BYLOC1 1.587 1.422 1.538 1.471 0.000
## BYLOC2 -5.280 4.250 0.741 -1.798 3.833
## BYLOC3 -0.384 -1.667 -0.261 -0.452 -0.419
## BYLOC4 -0.905 0.860 1.717 0.334 -2.196
##
##
## Normalized Residuals for Covariances/Correlations/Residual Correlations
## BYLOC2 BYLOC3 BYLOC4
## ________ ________ ________
## BYLOC2 0.000
## BYLOC3 -1.769 0.001
## BYLOC4 0.178 1.346 0.001
##
##
## ESTIMATED MODEL AND RESIDUALS (OBSERVED - ESTIMATED) FOR G2
##
##
## Model Estimated Covariances/Correlations/Residual Correlations
## BYCONF1 BYCONF2 BYCONF3 BYCONF4 BYLOC1
## ________ ________ ________ ________ ________
## BYCONF1 0.379
## BYCONF2 0.162 0.422
## BYCONF3 0.134 0.183 0.412
## BYCONF4 0.245 0.201 0.166 0.480
## BYLOC1 0.112 0.092 0.076 0.139 0.648
## BYLOC2 0.068 0.056 0.046 0.084 0.118
## BYLOC3 0.122 0.100 0.083 0.152 0.212
## BYLOC4 0.141 0.116 0.096 0.175 0.245
##
##
## Model Estimated Covariances/Correlations/Residual Correlations
## BYLOC2 BYLOC3 BYLOC4
## ________ ________ ________
## BYLOC2 0.494
## BYLOC3 0.128 0.568
## BYLOC4 0.148 0.268 0.634
##
##
## Residuals for Covariances/Correlations/Residual Correlations
## BYCONF1 BYCONF2 BYCONF3 BYCONF4 BYLOC1
## ________ ________ ________ ________ ________
## BYCONF1 0.000
## BYCONF2 0.005 0.000
## BYCONF3 0.000 0.000 0.000
## BYCONF4 0.003 -0.007 -0.004 0.000
## BYLOC1 -0.002 0.017 0.011 0.018 0.000
## BYLOC2 -0.035 0.017 0.009 -0.009 0.039
## BYLOC3 -0.014 -0.004 0.010 -0.005 -0.007
## BYLOC4 -0.009 0.013 0.014 0.008 -0.016
##
##
## Residuals for Covariances/Correlations/Residual Correlations
## BYLOC2 BYLOC3 BYLOC4
## ________ ________ ________
## BYLOC2 0.000
## BYLOC3 -0.006 0.000
## BYLOC4 -0.008 0.011 0.000
##
##
## Standardized Residuals (z-scores) for Covariances/Correlations/Residual Corr
## BYCONF1 BYCONF2 BYCONF3 BYCONF4 BYLOC1
## ________ ________ ________ ________ ________
## BYCONF1 0.000
## BYCONF2 2.764 0.069
## BYCONF3 0.181 0.000 0.000
## BYCONF4 3.411 -6.031 -2.643 0.000
## BYLOC1 -0.351 3.186 2.016 3.870 0.066
## BYLOC2 -8.072 3.225 1.658 -1.805 7.682
## BYLOC3 -3.892 -0.806 1.995 -1.344 -2.342
## BYLOC4 -2.690 2.671 2.750 2.381 -6.425
##
##
## Standardized Residuals (z-scores) for Covariances/Correlations/Residual Corr
## BYLOC2 BYLOC3 BYLOC4
## ________ ________ ________
## BYLOC2 0.000
## BYLOC3 -1.593 0.000
## BYLOC4 -2.383 5.255 0.000
##
##
## Normalized Residuals for Covariances/Correlations/Residual Correlations
## BYCONF1 BYCONF2 BYCONF3 BYCONF4 BYLOC1
## ________ ________ ________ ________ ________
## BYCONF1 0.000
## BYCONF2 0.900 0.000
## BYCONF3 0.071 0.000 0.000
## BYCONF4 0.399 -1.128 -0.705 0.000
## BYLOC1 -0.239 2.524 1.684 2.442 0.000
## BYLOC2 -6.299 2.786 1.486 -1.340 5.086
## BYLOC3 -2.305 -0.592 1.579 -0.706 -0.892
## BYLOC4 -1.446 1.877 2.095 1.119 -1.830
##
##
## Normalized Residuals for Covariances/Correlations/Residual Correlations
## BYLOC2 BYLOC3 BYLOC4
## ________ ________ ________
## BYLOC2 0.000
## BYLOC3 -0.853 0.000
## BYLOC4 -1.059 1.307 0.000
##
##
## Beginning Time: 12:19:51
## Ending Time: 12:19:51
## Elapsed Time: 00:00:00
##
##
##
## MUTHEN & MUTHEN
## 3463 Stoner Ave.
## Los Angeles, CA 90066
##
## Tel: (310) 391-9971
## Fax: (310) 391-8971
## Web: www.StatModel.com
## Support: Support@StatModel.com
##
## Copyright (c) 1998-2019 Muthen & Muthen6.2.2.2 constrain factor loadings (metric invariance)
TITLE: both groups, factor loading constraints
DATA: FILE IS "data\NELSTWOGROUPS.txt";
TYPE IS CORRELATION STDEVIATIONS;
NOBSERVATIONS ARE 5312 6008;
NGROUPS IS 2;
VARIABLE: NAMES ARE byconf1-byconf4 byloc1-byloc4;
ANALYSIS: ESTIMATOR=ML;
OUTPUT: SAMPSTAT STDYX MODINDICES(0);
MODEL:
f1 BY byconf1-byconf4;
f2 BY byloc1-byloc4;
f1 WITH f2;
byconf2 WITH byconf3;## Mplus VERSION 8.4
## MUTHEN & MUTHEN
## 06/10/2021 12:19 PM
##
## INPUT INSTRUCTIONS
##
## TITLE: both groups, factor loading constraints
## DATA: FILE IS "data\NELSTWOGROUPS.txt";
## TYPE IS CORRELATION STDEVIATIONS;
## NOBSERVATIONS ARE 5312 6008;
## NGROUPS IS 2;
## VARIABLE: NAMES ARE byconf1-byconf4 byloc1-byloc4;
## ANALYSIS: ESTIMATOR=ML;
## OUTPUT: SAMPSTAT STDYX MODINDICES(0);
## MODEL:
## f1 BY byconf1-byconf4;
## f2 BY byloc1-byloc4;
## f1 WITH f2;
## byconf2 WITH byconf3;
##
##
##
## 1 ERROR(S) FOUND IN THE INPUT INSTRUCTIONS
##
##
##
## both groups, factor loading constraints
##
## SUMMARY OF ANALYSIS
##
## Number of groups 2
## Number of observations
## Group G1 5312
## Group G2 6008
## Total sample size 11320
##
## Number of dependent variables 8
## Number of independent variables 0
## Number of continuous latent variables 2
##
## Observed dependent variables
##
## Continuous
## BYCONF1 BYCONF2 BYCONF3 BYCONF4 BYLOC1 BYLOC2
## BYLOC3 BYLOC4
##
## Continuous latent variables
## F1 F2
##
##
## Estimator ML
## Information matrix EXPECTED
## Maximum number of iterations 1000
## Convergence criterion 0.500D-04
## Maximum number of steepest descent iterations 20
##
## Input data file(s)
## data\NELSTWOGROUPS.txt
##
## Input data format FREE
##
##
## SAMPLE STATISTICS
##
##
## SAMPLE STATISTICS FOR G1
##
##
## Covariances/Correlations/Residual Correlations
## BYCONF1 BYCONF2 BYCONF3 BYCONF4 BYLOC1
## ________ ________ ________ ________ ________
## BYCONF1 0.343
## BYCONF2 0.125 0.413
## BYCONF3 0.101 0.166 0.384
## BYCONF4 0.184 0.155 0.126 0.426
## BYLOC1 0.096 0.084 0.070 0.119 0.646
## BYLOC2 0.025 0.077 0.045 0.060 0.146
## BYLOC3 0.093 0.071 0.065 0.118 0.185
## BYLOC4 0.095 0.093 0.082 0.130 0.180
##
##
## Covariances/Correlations/Residual Correlations
## BYLOC2 BYLOC3 BYLOC4
## ________ ________ ________
## BYLOC2 0.560
## BYLOC3 0.112 0.582
## BYLOC4 0.135 0.234 0.587
##
##
## SAMPLE STATISTICS FOR G2
##
##
## Covariances/Correlations/Residual Correlations
## BYCONF1 BYCONF2 BYCONF3 BYCONF4 BYLOC1
## ________ ________ ________ ________ ________
## BYCONF1 0.379
## BYCONF2 0.167 0.423
## BYCONF3 0.134 0.183 0.412
## BYCONF4 0.248 0.194 0.162 0.480
## BYLOC1 0.111 0.109 0.087 0.157 0.648
## BYLOC2 0.032 0.072 0.055 0.076 0.156
## BYLOC3 0.108 0.097 0.093 0.147 0.205
## BYLOC4 0.132 0.129 0.110 0.184 0.229
##
##
## Covariances/Correlations/Residual Correlations
## BYLOC2 BYLOC3 BYLOC4
## ________ ________ ________
## BYLOC2 0.494
## BYLOC3 0.122 0.569
## BYLOC4 0.140 0.279 0.634
##
##
## THE MODEL ESTIMATION TERMINATED NORMALLY
##
##
##
## MODEL FIT INFORMATION
##
## Number of Free Parameters 30
##
## Loglikelihood
##
## H0 Value -87848.453
## H1 Value -87671.417
##
## Information Criteria
##
## Akaike (AIC) 175756.907
## Bayesian (BIC) 175976.937
## Sample-Size Adjusted BIC 175881.600
## (n* = (n + 2) / 24)
##
## Chi-Square Test of Model Fit
##
## Value 354.073
## Degrees of Freedom 42
## P-Value 0.0000
##
## Chi-Square Contribution From Each Group
##
## G1 152.398
## G2 201.675
##
## RMSEA (Root Mean Square Error Of Approximation)
##
## Estimate 0.036
## 90 Percent C.I. 0.033 0.040
## Probability RMSEA <= .05 1.000
##
## CFI/TLI
##
## CFI 0.981
## TLI 0.975
##
## Chi-Square Test of Model Fit for the Baseline Model
##
## Value 16850.155
## Degrees of Freedom 56
## P-Value 0.0000
##
## SRMR (Standardized Root Mean Square Residual)
##
## Value 0.024
##
##
##
## MODEL RESULTS
##
## Two-Tailed
## Estimate S.E. Est./S.E. P-Value
##
## Group G1
##
## F1 BY
## BYCONF1 1.000 0.000 999.000 999.000
## BYCONF2 0.835 0.018 45.707 0.000
## BYCONF3 0.685 0.018 39.003 0.000
## BYCONF4 1.251 0.023 53.805 0.000
##
## F2 BY
## BYLOC1 1.000 0.000 999.000 999.000
## BYLOC2 0.628 0.022 28.972 0.000
## BYLOC3 1.101 0.028 39.508 0.000
## BYLOC4 1.228 0.031 40.165 0.000
##
## F1 WITH
## F2 0.086 0.004 22.131 0.000
##
## BYCONF2 WITH
## BYCONF3 0.080 0.005 16.041 0.000
##
## Variances
## F1 0.148 0.005 27.846 0.000
## F2 0.167 0.008 21.459 0.000
##
## Residual Variances
## BYCONF1 0.198 0.005 38.072 0.000
## BYCONF2 0.308 0.007 45.649 0.000
## BYCONF3 0.314 0.007 47.520 0.000
## BYCONF4 0.195 0.006 30.236 0.000
## BYLOC1 0.479 0.011 44.100 0.000
## BYLOC2 0.488 0.010 48.755 0.000
## BYLOC3 0.375 0.009 39.722 0.000
## BYLOC4 0.342 0.010 35.161 0.000
##
## Group G2
##
## F1 BY
## BYCONF1 1.000 0.000 999.000 999.000
## BYCONF2 0.835 0.018 45.707 0.000
## BYCONF3 0.685 0.018 39.003 0.000
## BYCONF4 1.251 0.023 53.805 0.000
##
## F2 BY
## BYLOC1 1.000 0.000 999.000 999.000
## BYLOC2 0.628 0.022 28.972 0.000
## BYLOC3 1.101 0.028 39.508 0.000
## BYLOC4 1.228 0.031 40.165 0.000
##
## F1 WITH
## F2 0.112 0.004 25.256 0.000
##
## BYCONF2 WITH
## BYCONF3 0.073 0.005 15.777 0.000
##
## Variances
## F1 0.195 0.006 30.644 0.000
## F2 0.196 0.009 22.735 0.000
##
## Residual Variances
## BYCONF1 0.183 0.005 38.038 0.000
## BYCONF2 0.289 0.006 47.749 0.000
## BYCONF3 0.321 0.006 50.417 0.000
## BYCONF4 0.175 0.006 28.603 0.000
## BYLOC1 0.452 0.010 46.176 0.000
## BYLOC2 0.421 0.008 51.230 0.000
## BYLOC3 0.334 0.008 40.210 0.000
## BYLOC4 0.331 0.009 36.475 0.000
##
##
## STANDARDIZED MODEL RESULTS
##
##
## STDYX Standardization
##
## Two-Tailed
## Estimate S.E. Est./S.E. P-Value
##
## Group G1
##
## F1 BY
## BYCONF1 0.654 0.009 70.075 0.000
## BYCONF2 0.501 0.010 51.537 0.000
## BYCONF3 0.425 0.010 42.270 0.000
## BYCONF4 0.736 0.009 78.543 0.000
##
## F2 BY
## BYLOC1 0.509 0.011 48.036 0.000
## BYLOC2 0.345 0.011 32.845 0.000
## BYLOC3 0.592 0.011 56.183 0.000
## BYLOC4 0.652 0.011 61.318 0.000
##
## F1 WITH
## F2 0.545 0.017 31.967 0.000
##
## BYCONF2 WITH
## BYCONF3 0.257 0.014 18.373 0.000
##
## Variances
## F1 1.000 0.000 999.000 999.000
## F2 1.000 0.000 999.000 999.000
##
## Residual Variances
## BYCONF1 0.572 0.012 46.918 0.000
## BYCONF2 0.749 0.010 77.038 0.000
## BYCONF3 0.819 0.009 95.754 0.000
## BYCONF4 0.458 0.014 33.130 0.000
## BYLOC1 0.741 0.011 68.822 0.000
## BYLOC2 0.881 0.007 121.324 0.000
## BYLOC3 0.649 0.012 52.049 0.000
## BYLOC4 0.575 0.014 41.542 0.000
##
## Group G2
##
## F1 BY
## BYCONF1 0.719 0.008 86.377 0.000
## BYCONF2 0.566 0.010 59.562 0.000
## BYCONF3 0.471 0.010 46.264 0.000
## BYCONF4 0.797 0.008 100.044 0.000
##
## F2 BY
## BYLOC1 0.550 0.010 53.518 0.000
## BYLOC2 0.394 0.011 35.315 0.000
## BYLOC3 0.645 0.010 65.444 0.000
## BYLOC4 0.687 0.010 70.837 0.000
##
## F1 WITH
## F2 0.571 0.014 41.122 0.000
##
## BYCONF2 WITH
## BYCONF3 0.239 0.013 17.852 0.000
##
## Variances
## F1 1.000 0.000 999.000 999.000
## F2 1.000 0.000 999.000 999.000
##
## Residual Variances
## BYCONF1 0.483 0.012 40.425 0.000
## BYCONF2 0.680 0.011 63.199 0.000
## BYCONF3 0.778 0.010 81.055 0.000
## BYCONF4 0.365 0.013 28.748 0.000
## BYLOC1 0.697 0.011 61.607 0.000
## BYLOC2 0.845 0.009 95.949 0.000
## BYLOC3 0.584 0.013 45.966 0.000
## BYLOC4 0.528 0.013 39.641 0.000
##
##
## R-SQUARE
##
## Group G1
##
## Observed Two-Tailed
## Variable Estimate S.E. Est./S.E. P-Value
##
## BYCONF1 0.428 0.012 35.037 0.000
## BYCONF2 0.251 0.010 25.769 0.000
## BYCONF3 0.181 0.009 21.135 0.000
## BYCONF4 0.542 0.014 39.272 0.000
## BYLOC1 0.259 0.011 24.018 0.000
## BYLOC2 0.119 0.007 16.423 0.000
## BYLOC3 0.351 0.012 28.091 0.000
## BYLOC4 0.425 0.014 30.659 0.000
##
## Group G2
##
## Observed Two-Tailed
## Variable Estimate S.E. Est./S.E. P-Value
##
## BYCONF1 0.517 0.012 43.189 0.000
## BYCONF2 0.320 0.011 29.781 0.000
## BYCONF3 0.222 0.010 23.132 0.000
## BYCONF4 0.635 0.013 50.022 0.000
## BYLOC1 0.303 0.011 26.759 0.000
## BYLOC2 0.155 0.009 17.658 0.000
## BYLOC3 0.416 0.013 32.722 0.000
## BYLOC4 0.472 0.013 35.419 0.000
##
##
## QUALITY OF NUMERICAL RESULTS
##
## Condition Number for the Information Matrix 0.323E-02
## (ratio of smallest to largest eigenvalue)
##
##
## MODEL MODIFICATION INDICES
##
## NOTE: Modification indices for direct effects of observed dependent variables
## regressed on covariates may not be included. To include these, request
## MODINDICES (ALL).
##
## Minimum M.I. value for printing the modification index 0.000
##
## M.I. E.P.C. Std E.P.C. StdYX E.P.C.
## Group G1
##
##
## BY Statements
##
## F1 BY BYCONF1 0.989 -0.035 -0.013 -0.023
## F1 BY BYCONF2 0.690 0.017 0.006 0.010
## F1 BY BYCONF3 0.003 0.001 0.000 0.001
## F1 BY BYCONF4 0.036 0.005 0.002 0.003
## F1 BY BYLOC1 7.401 0.099 0.038 0.047
## F1 BY BYLOC2 1.455 -0.041 -0.016 -0.021
## F1 BY BYLOC3 0.170 -0.014 -0.006 -0.007
## F1 BY BYLOC4 1.388 -0.043 -0.017 -0.022
## F2 BY BYCONF1 5.210 -0.057 -0.023 -0.040
## F2 BY BYCONF2 1.531 0.032 0.013 0.020
## F2 BY BYCONF3 1.643 0.032 0.013 0.021
## F2 BY BYCONF4 0.150 0.011 0.005 0.007
## F2 BY BYLOC1 0.001 -0.002 -0.001 -0.001
## F2 BY BYLOC2 3.540 0.046 0.019 0.025
## F2 BY BYLOC3 1.378 0.033 0.013 0.018
## F2 BY BYLOC4 5.467 -0.072 -0.029 -0.038
##
## WITH Statements
##
## BYCONF2 WITH BYCONF1 0.103 0.001 0.001 0.005
## BYCONF3 WITH BYCONF1 0.042 -0.001 -0.001 -0.003
## BYCONF4 WITH BYCONF1 0.547 0.005 0.005 0.024
## BYCONF4 WITH BYCONF2 0.056 -0.001 -0.001 -0.005
## BYCONF4 WITH BYCONF3 0.467 -0.003 -0.003 -0.013
## BYLOC1 WITH BYCONF1 3.608 0.010 0.010 0.032
## BYLOC1 WITH BYCONF2 0.224 0.003 0.003 0.007
## BYLOC1 WITH BYCONF3 0.260 0.003 0.003 0.007
## BYLOC1 WITH BYCONF4 0.953 0.005 0.005 0.018
## BYLOC2 WITH BYCONF1 40.514 -0.031 -0.031 -0.101
## BYLOC2 WITH BYCONF2 40.885 0.035 0.035 0.090
## BYLOC2 WITH BYCONF3 0.043 -0.001 -0.001 -0.003
## BYLOC2 WITH BYCONF4 1.091 -0.006 -0.006 -0.018
## BYLOC2 WITH BYLOC1 37.664 0.046 0.046 0.095
## BYLOC3 WITH BYCONF1 0.897 0.005 0.005 0.017
## BYLOC3 WITH BYCONF2 7.153 -0.014 -0.014 -0.041
## BYLOC3 WITH BYCONF3 0.012 -0.001 -0.001 -0.002
## BYLOC3 WITH BYCONF4 0.001 0.000 0.000 0.001
## BYLOC3 WITH BYLOC1 0.019 -0.001 -0.001 -0.003
## BYLOC3 WITH BYLOC2 4.043 -0.014 -0.014 -0.033
## BYLOC4 WITH BYCONF1 2.638 -0.008 -0.008 -0.030
## BYLOC4 WITH BYCONF2 0.013 0.001 0.001 0.002
## BYLOC4 WITH BYCONF3 2.491 0.008 0.008 0.025
## BYLOC4 WITH BYCONF4 0.021 0.001 0.001 0.003
## BYLOC4 WITH BYLOC1 35.153 -0.050 -0.050 -0.123
## BYLOC4 WITH BYLOC2 0.219 0.003 0.003 0.008
## BYLOC4 WITH BYLOC3 11.347 0.029 0.029 0.082
##
## Group G2
##
##
## BY Statements
##
## F1 BY BYCONF1 0.994 0.035 0.015 0.025
## F1 BY BYCONF2 0.690 -0.009 -0.004 -0.006
## F1 BY BYCONF3 0.003 -0.001 0.000 0.000
## F1 BY BYCONF4 0.037 -0.003 -0.001 -0.002
## F1 BY BYLOC1 7.753 0.084 0.037 0.046
## F1 BY BYLOC2 16.415 -0.109 -0.048 -0.068
## F1 BY BYLOC3 8.422 -0.084 -0.037 -0.049
## F1 BY BYLOC4 8.089 0.088 0.039 0.049
## F2 BY BYCONF1 24.068 -0.111 -0.049 -0.080
## F2 BY BYCONF2 2.204 0.034 0.015 0.023
## F2 BY BYCONF3 5.676 0.055 0.025 0.038
## F2 BY BYCONF4 3.912 0.053 0.024 0.034
## F2 BY BYLOC1 0.001 0.001 0.001 0.001
## F2 BY BYLOC2 3.538 -0.028 -0.012 -0.017
## F2 BY BYLOC3 1.366 -0.020 -0.009 -0.012
## F2 BY BYLOC4 5.476 0.045 0.020 0.025
##
## WITH Statements
##
## BYCONF2 WITH BYCONF1 5.726 0.010 0.010 0.042
## BYCONF3 WITH BYCONF1 0.088 -0.001 -0.001 -0.005
## BYCONF4 WITH BYCONF1 15.235 0.027 0.027 0.149
## BYCONF4 WITH BYCONF2 16.951 -0.020 -0.020 -0.088
## BYCONF4 WITH BYCONF3 1.922 -0.006 -0.006 -0.027
## BYLOC1 WITH BYCONF1 0.283 -0.002 -0.002 -0.008
## BYLOC1 WITH BYCONF2 2.121 0.007 0.007 0.020
## BYLOC1 WITH BYCONF3 0.166 -0.002 -0.002 -0.006
## BYLOC1 WITH BYCONF4 6.372 0.013 0.013 0.045
## BYLOC2 WITH BYCONF1 57.623 -0.032 -0.032 -0.115
## BYLOC2 WITH BYCONF2 18.094 0.020 0.020 0.057
## BYLOC2 WITH BYCONF3 1.191 0.005 0.005 0.014
## BYLOC2 WITH BYCONF4 0.035 -0.001 -0.001 -0.003
## BYLOC2 WITH BYLOC1 55.123 0.048 0.048 0.110
## BYLOC3 WITH BYCONF1 0.702 -0.003 -0.003 -0.014
## BYLOC3 WITH BYCONF2 5.208 -0.010 -0.010 -0.033
## BYLOC3 WITH BYCONF3 3.918 0.009 0.009 0.028
## BYLOC3 WITH BYCONF4 1.393 -0.005 -0.005 -0.022
## BYLOC3 WITH BYLOC1 7.483 -0.020 -0.020 -0.051
## BYLOC3 WITH BYLOC2 5.062 -0.014 -0.014 -0.036
## BYLOC4 WITH BYCONF1 1.047 -0.004 -0.004 -0.018
## BYLOC4 WITH BYCONF2 1.252 0.005 0.005 0.017
## BYLOC4 WITH BYCONF3 0.992 0.005 0.005 0.015
## BYLOC4 WITH BYCONF4 2.221 0.007 0.007 0.030
## BYLOC4 WITH BYLOC1 20.294 -0.035 -0.035 -0.092
## BYLOC4 WITH BYLOC2 5.493 -0.015 -0.015 -0.040
## BYLOC4 WITH BYLOC3 32.803 0.047 0.047 0.143
##
##
##
## Beginning Time: 12:19:51
## Ending Time: 12:19:52
## Elapsed Time: 00:00:01
##
##
##
## MUTHEN & MUTHEN
## 3463 Stoner Ave.
## Los Angeles, CA 90066
##
## Tel: (310) 391-9971
## Fax: (310) 391-8971
## Web: www.StatModel.com
## Support: Support@StatModel.com
##
## Copyright (c) 1998-2019 Muthen & Muthen6.2.2.3 release constraints across groups
Use modification indices and theory as guidelines
TITLE: both groups, factor loading constraints
DATA: FILE IS "data\NELSTWOGROUPS.txt";
TYPE IS CORRELATION STDEVIATIONS;
NOBSERVATIONS ARE 5312 6008;
NGROUPS IS 2;
VARIABLE: NAMES ARE byconf1-byconf4 byloc1-byloc4;
ANALYSIS: ESTIMATOR=ML;
OUTPUT: SAMPSTAT STDYX MODINDICES(0);
MODEL:
f1 BY byconf1-byconf4;
f2 BY byloc1-byloc4;
f1 WITH f2;
byconf2 WITH byconf3;
MODEL g2:
f2 BY loc2*;## Mplus VERSION 8.4
## MUTHEN & MUTHEN
## 06/10/2021 12:19 PM
##
## INPUT INSTRUCTIONS
##
## TITLE: both groups, factor loading constraints
## DATA: FILE IS "data\NELSTWOGROUPS.txt";
## TYPE IS CORRELATION STDEVIATIONS;
## NOBSERVATIONS ARE 5312 6008;
## NGROUPS IS 2;
## VARIABLE: NAMES ARE byconf1-byconf4 byloc1-byloc4;
## ANALYSIS: ESTIMATOR=ML;
## OUTPUT: SAMPSTAT STDYX MODINDICES(0);
## MODEL:
## f1 BY byconf1-byconf4;
## f2 BY byloc1-byloc4;
## f1 WITH f2;
## byconf2 WITH byconf3;
## MODEL g2:
## f2 BY loc2*;
##
## *** ERROR in MODEL command
## Unknown variable(s) in a BY statement: LOC2
##
##
## MUTHEN & MUTHEN
## 3463 Stoner Ave.
## Los Angeles, CA 90066
##
## Tel: (310) 391-9971
## Fax: (310) 391-8971
## Web: www.StatModel.com
## Support: Support@StatModel.com
##
## Copyright (c) 1998-2019 Muthen & Muthen6.2.2.4 constrain loadings, residual variances, and residual covariances
TITLE: both groups, factor loading and
residual variances and covariances constraints
DATA: FILE IS "data\NELSTWOGROUPS.txt";
TYPE IS CORRELATION STDEVIATIONS;
NOBSERVATIONS ARE 5312 6008;
NGROUPS IS 2;
VARIABLE: NAMES ARE byconf1-byconf4 byloc1-byloc4;
ANALYSIS: ESTIMATOR=ML;
OUTPUT: STDYX;
MODEL:
f1 BY byconf1-byconf4;
f2 BY byloc1-byloc4;
f1 WITH f2;
byconf1 (1);
byconf2 (2);
byconf3 (3);
byconf4 (4);
byloc1 (5);
byloc2 (6);
byloc3 (7);
byloc4 (8);
byconf2 WITH byconf3 (9);## Mplus VERSION 8.4
## MUTHEN & MUTHEN
## 06/10/2021 12:19 PM
##
## INPUT INSTRUCTIONS
##
## TITLE: both groups, factor loading and
## residual variances and covariances constraints
## DATA: FILE IS "data\NELSTWOGROUPS.txt";
## TYPE IS CORRELATION STDEVIATIONS;
## NOBSERVATIONS ARE 5312 6008;
## NGROUPS IS 2;
## VARIABLE: NAMES ARE byconf1-byconf4 byloc1-byloc4;
## ANALYSIS: ESTIMATOR=ML;
## OUTPUT: STDYX;
## MODEL:
## f1 BY byconf1-byconf4;
## f2 BY byloc1-byloc4;
## f1 WITH f2;
## byconf1 (1);
## byconf2 (2);
## byconf3 (3);
## byconf4 (4);
## byloc1 (5);
## byloc2 (6);
## byloc3 (7);
## byloc4 (8);
## byconf2 WITH byconf3 (9);
##
##
##
## 1 ERROR(S) FOUND IN THE INPUT INSTRUCTIONS
##
##
##
## both groups, factor loading and
## residual variances and covariances constraints
##
## SUMMARY OF ANALYSIS
##
## Number of groups 2
## Number of observations
## Group G1 5312
## Group G2 6008
## Total sample size 11320
##
## Number of dependent variables 8
## Number of independent variables 0
## Number of continuous latent variables 2
##
## Observed dependent variables
##
## Continuous
## BYCONF1 BYCONF2 BYCONF3 BYCONF4 BYLOC1 BYLOC2
## BYLOC3 BYLOC4
##
## Continuous latent variables
## F1 F2
##
##
## Estimator ML
## Information matrix EXPECTED
## Maximum number of iterations 1000
## Convergence criterion 0.500D-04
## Maximum number of steepest descent iterations 20
##
## Input data file(s)
## data\NELSTWOGROUPS.txt
##
## Input data format FREE
##
##
##
## THE MODEL ESTIMATION TERMINATED NORMALLY
##
##
##
## MODEL FIT INFORMATION
##
## Number of Free Parameters 21
##
## Loglikelihood
##
## H0 Value -87881.275
## H1 Value -87671.417
##
## Information Criteria
##
## Akaike (AIC) 175804.549
## Bayesian (BIC) 175958.570
## Sample-Size Adjusted BIC 175891.835
## (n* = (n + 2) / 24)
##
## Chi-Square Test of Model Fit
##
## Value 419.715
## Degrees of Freedom 51
## P-Value 0.0000
##
## Chi-Square Contribution From Each Group
##
## G1 185.577
## G2 234.138
##
## RMSEA (Root Mean Square Error Of Approximation)
##
## Estimate 0.036
## 90 Percent C.I. 0.033 0.039
## Probability RMSEA <= .05 1.000
##
## CFI/TLI
##
## CFI 0.978
## TLI 0.976
##
## Chi-Square Test of Model Fit for the Baseline Model
##
## Value 16850.155
## Degrees of Freedom 56
## P-Value 0.0000
##
## SRMR (Standardized Root Mean Square Residual)
##
## Value 0.028
##
##
##
## MODEL RESULTS
##
## Two-Tailed
## Estimate S.E. Est./S.E. P-Value
##
## Group G1
##
## F1 BY
## BYCONF1 1.000 0.000 999.000 999.000
## BYCONF2 0.837 0.018 45.507 0.000
## BYCONF3 0.687 0.018 38.969 0.000
## BYCONF4 1.252 0.023 53.506 0.000
##
## F2 BY
## BYLOC1 1.000 0.000 999.000 999.000
## BYLOC2 0.631 0.022 28.887 0.000
## BYLOC3 1.101 0.028 39.370 0.000
## BYLOC4 1.227 0.031 40.049 0.000
##
## F1 WITH
## F2 0.086 0.004 22.086 0.000
##
## BYCONF2 WITH
## BYCONF3 0.076 0.003 21.938 0.000
##
## Variances
## F1 0.150 0.005 28.430 0.000
## F2 0.171 0.008 21.772 0.000
##
## Residual Variances
## BYCONF1 0.190 0.004 50.559 0.000
## BYCONF2 0.297 0.005 64.604 0.000
## BYCONF3 0.318 0.005 68.309 0.000
## BYCONF4 0.185 0.005 37.663 0.000
## BYLOC1 0.464 0.008 61.888 0.000
## BYLOC2 0.452 0.006 69.908 0.000
## BYLOC3 0.353 0.007 53.462 0.000
## BYLOC4 0.336 0.007 47.135 0.000
##
## Group G2
##
## F1 BY
## BYCONF1 1.000 0.000 999.000 999.000
## BYCONF2 0.837 0.018 45.507 0.000
## BYCONF3 0.687 0.018 38.969 0.000
## BYCONF4 1.252 0.023 53.506 0.000
##
## F2 BY
## BYLOC1 1.000 0.000 999.000 999.000
## BYLOC2 0.631 0.022 28.887 0.000
## BYLOC3 1.101 0.028 39.370 0.000
## BYLOC4 1.227 0.031 40.049 0.000
##
## F1 WITH
## F2 0.112 0.004 25.265 0.000
##
## BYCONF2 WITH
## BYCONF3 0.076 0.003 21.938 0.000
##
## Variances
## F1 0.192 0.006 30.451 0.000
## F2 0.193 0.009 22.666 0.000
##
## Residual Variances
## BYCONF1 0.190 0.004 50.559 0.000
## BYCONF2 0.297 0.005 64.604 0.000
## BYCONF3 0.318 0.005 68.309 0.000
## BYCONF4 0.185 0.005 37.663 0.000
## BYLOC1 0.464 0.008 61.888 0.000
## BYLOC2 0.452 0.006 69.908 0.000
## BYLOC3 0.353 0.007 53.462 0.000
## BYLOC4 0.336 0.007 47.135 0.000
##
##
## STANDARDIZED MODEL RESULTS
##
##
## STDYX Standardization
##
## Two-Tailed
## Estimate S.E. Est./S.E. P-Value
##
## Group G1
##
## F1 BY
## BYCONF1 0.665 0.009 78.075 0.000
## BYCONF2 0.511 0.009 55.397 0.000
## BYCONF3 0.427 0.010 44.293 0.000
## BYCONF4 0.749 0.008 90.140 0.000
##
## F2 BY
## BYLOC1 0.519 0.010 51.689 0.000
## BYLOC2 0.362 0.011 34.315 0.000
## BYLOC3 0.608 0.010 62.504 0.000
## BYLOC4 0.658 0.010 68.656 0.000
##
## F1 WITH
## F2 0.534 0.017 32.111 0.000
##
## BYCONF2 WITH
## BYCONF3 0.247 0.010 25.101 0.000
##
## Variances
## F1 1.000 0.000 999.000 999.000
## F2 1.000 0.000 999.000 999.000
##
## Residual Variances
## BYCONF1 0.558 0.011 49.370 0.000
## BYCONF2 0.739 0.009 78.327 0.000
## BYCONF3 0.818 0.008 99.345 0.000
## BYCONF4 0.440 0.012 35.349 0.000
## BYLOC1 0.731 0.010 70.214 0.000
## BYLOC2 0.869 0.008 114.028 0.000
## BYLOC3 0.630 0.012 53.238 0.000
## BYLOC4 0.567 0.013 44.872 0.000
##
## Group G2
##
## F1 BY
## BYCONF1 0.709 0.008 91.388 0.000
## BYCONF2 0.558 0.009 61.527 0.000
## BYCONF3 0.471 0.010 47.808 0.000
## BYCONF4 0.787 0.007 109.508 0.000
##
## F2 BY
## BYLOC1 0.542 0.010 55.152 0.000
## BYLOC2 0.381 0.011 35.540 0.000
## BYLOC3 0.631 0.009 67.851 0.000
## BYLOC4 0.681 0.009 75.316 0.000
##
## F1 WITH
## F2 0.581 0.014 41.515 0.000
##
## BYCONF2 WITH
## BYCONF3 0.247 0.010 25.101 0.000
##
## Variances
## F1 1.000 0.000 999.000 999.000
## F2 1.000 0.000 999.000 999.000
##
## Residual Variances
## BYCONF1 0.497 0.011 45.133 0.000
## BYCONF2 0.688 0.010 67.944 0.000
## BYCONF3 0.778 0.009 83.749 0.000
## BYCONF4 0.380 0.011 33.540 0.000
## BYLOC1 0.706 0.011 66.362 0.000
## BYLOC2 0.855 0.008 104.611 0.000
## BYLOC3 0.601 0.012 51.192 0.000
## BYLOC4 0.537 0.012 43.604 0.000
##
##
## R-SQUARE
##
## Group G1
##
## Observed Two-Tailed
## Variable Estimate S.E. Est./S.E. P-Value
##
## BYCONF1 0.442 0.011 39.038 0.000
## BYCONF2 0.261 0.009 27.698 0.000
## BYCONF3 0.182 0.008 22.146 0.000
## BYCONF4 0.560 0.012 45.070 0.000
## BYLOC1 0.269 0.010 25.844 0.000
## BYLOC2 0.131 0.008 17.158 0.000
## BYLOC3 0.370 0.012 31.252 0.000
## BYLOC4 0.433 0.013 34.328 0.000
##
## Group G2
##
## Observed Two-Tailed
## Variable Estimate S.E. Est./S.E. P-Value
##
## BYCONF1 0.503 0.011 45.694 0.000
## BYCONF2 0.312 0.010 30.764 0.000
## BYCONF3 0.222 0.009 23.904 0.000
## BYCONF4 0.620 0.011 54.754 0.000
## BYLOC1 0.294 0.011 27.576 0.000
## BYLOC2 0.145 0.008 17.770 0.000
## BYLOC3 0.399 0.012 33.926 0.000
## BYLOC4 0.463 0.012 37.658 0.000
##
##
## QUALITY OF NUMERICAL RESULTS
##
## Condition Number for the Information Matrix 0.318E-02
## (ratio of smallest to largest eigenvalue)
##
##
## Beginning Time: 12:19:52
## Ending Time: 12:19:52
## Elapsed Time: 00:00:00
##
##
##
## MUTHEN & MUTHEN
## 3463 Stoner Ave.
## Los Angeles, CA 90066
##
## Tel: (310) 391-9971
## Fax: (310) 391-8971
## Web: www.StatModel.com
## Support: Support@StatModel.com
##
## Copyright (c) 1998-2019 Muthen & Muthen6.2.3 CFA with Ordered Categorical Variables – An example from Wang & Su (2013)
TITLE: An example from Wang & Su (2013)
DATA: FILE IS "data\NELSSelfConcept3waves.dat";
FORMAT IS F7 F4 2F2 F11.4 39F2;
VARIABLE: NAMES ARE id status race gender weight
byconf1 byconf2 byconf3 byconf4 byconf5 byconf6 byconf7
byloc1 byloc2 byloc3 byloc4 byloc5 byloc6
f1conf1 f1conf2 f1conf3 f1conf4 f1conf5 f1conf6 f1conf7
f1loc1 f1loc2 f1loc3 f1loc4 f1loc5 f1loc6
f2conf1 f2conf2 f2conf3 f2conf4 f2conf5 f2conf6 f2conf7
f2loc1 f2loc2 f2loc3 f2loc4 f2loc5 f2loc6;
USEVARIABLES ARE byconf1-byconf4 byloc1-byloc4
f1conf1-f1conf4 f1loc1-f1loc4
f2conf1-f2conf4 f2loc1-f2loc4;
CATEGORICAL ARE byconf1-byconf4 byloc1-byloc4
f1conf1-f1conf4 f1loc1-f1loc4
f2conf1-f2conf4 f2loc1-f2loc4;
WEIGHT IS weight;
MISSING ARE race (99) gender (99) byconf1-f2loc6 (99);
ANALYSIS: ESTIMATOR=WLSMV;
OUTPUT: SAMPSTAT STANDARDIZED MODINDICES;
MODEL: f1 BY byconf1-byconf4;
f2 BY byloc1-byloc4;
f3 BY f1conf1-f1conf4;
f4 BY f1loc1-f1loc4;
f5 BY f2conf1-f2conf4;
f6 BY f2loc1-f2loc4;## Mplus VERSION 8.4
## MUTHEN & MUTHEN
## 06/10/2021 12:19 PM
##
## INPUT INSTRUCTIONS
##
## TITLE: An example from Wang & Su (2013)
## DATA: FILE IS "data\NELSSelfConcept3waves.dat";
## FORMAT IS F7 F4 2F2 F11.4 39F2;
## VARIABLE: NAMES ARE id status race gender weight
## byconf1 byconf2 byconf3 byconf4 byconf5 byconf6 byconf7
## byloc1 byloc2 byloc3 byloc4 byloc5 byloc6
## f1conf1 f1conf2 f1conf3 f1conf4 f1conf5 f1conf6 f1conf7
## f1loc1 f1loc2 f1loc3 f1loc4 f1loc5 f1loc6
## f2conf1 f2conf2 f2conf3 f2conf4 f2conf5 f2conf6 f2conf7
## f2loc1 f2loc2 f2loc3 f2loc4 f2loc5 f2loc6;
## USEVARIABLES ARE byconf1-byconf4 byloc1-byloc4
## f1conf1-f1conf4 f1loc1-f1loc4
## f2conf1-f2conf4 f2loc1-f2loc4;
## CATEGORICAL ARE byconf1-byconf4 byloc1-byloc4
## f1conf1-f1conf4 f1loc1-f1loc4
## f2conf1-f2conf4 f2loc1-f2loc4;
## WEIGHT IS weight;
## MISSING ARE race (99) gender (99) byconf1-f2loc6 (99);
## ANALYSIS: ESTIMATOR=WLSMV;
## OUTPUT: SAMPSTAT STANDARDIZED MODINDICES;
## MODEL: f1 BY byconf1-byconf4;
## f2 BY byloc1-byloc4;
## f3 BY f1conf1-f1conf4;
## f4 BY f1loc1-f1loc4;
## f5 BY f2conf1-f2conf4;
## f6 BY f2loc1-f2loc4;
##
##
##
## *** WARNING
## Data set contains cases with missing on all variables.
## These cases were not included in the analysis.
## Number of cases with missing on all variables: 155
## 1 ERROR(S) FOUND IN THE INPUT INSTRUCTIONS
## 1 WARNING(S) FOUND IN THE INPUT INSTRUCTIONS
##
##
##
## An example from Wang & Su (2013)
##
## SUMMARY OF ANALYSIS
##
## Number of groups 1
## Number of observations 11989
##
## Number of dependent variables 24
## Number of independent variables 0
## Number of continuous latent variables 6
##
## Observed dependent variables
##
## Binary and ordered categorical (ordinal)
## BYCONF1 BYCONF2 BYCONF3 BYCONF4 BYLOC1 BYLOC2
## BYLOC3 BYLOC4 F1CONF1 F1CONF2 F1CONF3 F1CONF4
## F1LOC1 F1LOC2 F1LOC3 F1LOC4 F2CONF1 F2CONF2
## F2CONF3 F2CONF4 F2LOC1 F2LOC2 F2LOC3 F2LOC4
##
## Continuous latent variables
## F1 F2 F3 F4 F5 F6
##
## Variables with special functions
##
## Weight variable WEIGHT
##
## Estimator WLSMV
## Maximum number of iterations 1000
## Convergence criterion 0.500D-04
## Maximum number of steepest descent iterations 20
## Maximum number of iterations for H1 2000
## Convergence criterion for H1 0.100D-03
## Parameterization DELTA
## Link PROBIT
##
## Input data file(s)
## data\NELSSelfConcept3waves.dat
##
## Input data format
## (F7 F4 2F2 F11.4 39F2)
##
##
## SUMMARY OF DATA
##
## Number of missing data patterns 273
##
##
## COVARIANCE COVERAGE OF DATA
##
## Minimum covariance coverage value 0.100
##
##
## PROPORTION OF DATA PRESENT
##
##
## Covariance Coverage
## BYCONF1 BYCONF2 BYCONF3 BYCONF4 BYLOC1
## ________ ________ ________ ________ ________
## BYCONF1 0.942
## BYCONF2 0.929 0.931
## BYCONF3 0.933 0.923 0.935
## BYCONF4 0.932 0.923 0.927 0.934
## BYLOC1 0.938 0.928 0.932 0.931 0.940
## BYLOC2 0.936 0.926 0.930 0.929 0.935
## BYLOC3 0.936 0.926 0.931 0.930 0.935
## BYLOC4 0.936 0.927 0.931 0.930 0.935
## F1CONF1 0.843 0.834 0.838 0.837 0.842
## F1CONF2 0.840 0.831 0.835 0.834 0.839
## F1CONF3 0.836 0.827 0.831 0.830 0.835
## F1CONF4 0.837 0.829 0.832 0.832 0.836
## F1LOC1 0.840 0.831 0.834 0.834 0.838
## F1LOC2 0.838 0.830 0.833 0.833 0.837
## F1LOC3 0.839 0.830 0.834 0.833 0.838
## F1LOC4 0.836 0.827 0.831 0.831 0.835
## F2CONF1 0.780 0.771 0.775 0.774 0.779
## F2CONF2 0.773 0.764 0.767 0.767 0.772
## F2CONF3 0.773 0.765 0.767 0.767 0.772
## F2CONF4 0.774 0.765 0.768 0.768 0.773
## F2LOC1 0.777 0.769 0.772 0.771 0.776
## F2LOC2 0.776 0.768 0.771 0.771 0.775
## F2LOC3 0.775 0.766 0.769 0.769 0.774
## F2LOC4 0.772 0.764 0.767 0.766 0.771
##
##
## Covariance Coverage
## BYLOC2 BYLOC3 BYLOC4 F1CONF1 F1CONF2
## ________ ________ ________ ________ ________
## BYLOC2 0.938
## BYLOC3 0.933 0.938
## BYLOC4 0.933 0.934 0.938
## F1CONF1 0.840 0.840 0.841 0.885
## F1CONF2 0.837 0.837 0.838 0.881 0.882
## F1CONF3 0.833 0.833 0.834 0.877 0.875
## F1CONF4 0.834 0.834 0.835 0.878 0.876
## F1LOC1 0.836 0.837 0.837 0.881 0.878
## F1LOC2 0.835 0.835 0.836 0.880 0.877
## F1LOC3 0.836 0.836 0.836 0.880 0.878
## F1LOC4 0.833 0.833 0.834 0.877 0.875
## F2CONF1 0.777 0.777 0.777 0.761 0.758
## F2CONF2 0.770 0.770 0.770 0.755 0.752
## F2CONF3 0.770 0.770 0.771 0.755 0.752
## F2CONF4 0.771 0.770 0.771 0.756 0.753
## F2LOC1 0.774 0.774 0.775 0.759 0.756
## F2LOC2 0.773 0.773 0.774 0.758 0.755
## F2LOC3 0.772 0.772 0.772 0.757 0.754
## F2LOC4 0.769 0.769 0.770 0.754 0.751
##
##
## Covariance Coverage
## F1CONF3 F1CONF4 F1LOC1 F1LOC2 F1LOC3
## ________ ________ ________ ________ ________
## F1CONF3 0.878
## F1CONF4 0.873 0.879
## F1LOC1 0.874 0.875 0.882
## F1LOC2 0.873 0.874 0.877 0.880
## F1LOC3 0.875 0.876 0.877 0.876 0.881
## F1LOC4 0.872 0.874 0.874 0.873 0.875
## F2CONF1 0.755 0.756 0.758 0.757 0.758
## F2CONF2 0.749 0.750 0.751 0.751 0.752
## F2CONF3 0.749 0.750 0.752 0.751 0.752
## F2CONF4 0.750 0.751 0.752 0.752 0.752
## F2LOC1 0.753 0.754 0.755 0.755 0.755
## F2LOC2 0.752 0.753 0.754 0.754 0.754
## F2LOC3 0.751 0.752 0.753 0.753 0.753
## F2LOC4 0.748 0.749 0.751 0.750 0.751
##
##
## Covariance Coverage
## F1LOC4 F2CONF1 F2CONF2 F2CONF3 F2CONF4
## ________ ________ ________ ________ ________
## F1LOC4 0.877
## F2CONF1 0.755 0.823
## F2CONF2 0.749 0.815 0.815
## F2CONF3 0.749 0.814 0.810 0.815
## F2CONF4 0.750 0.816 0.811 0.811 0.816
## F2LOC1 0.753 0.819 0.812 0.812 0.813
## F2LOC2 0.752 0.818 0.812 0.812 0.813
## F2LOC3 0.751 0.817 0.812 0.813 0.814
## F2LOC4 0.748 0.814 0.809 0.810 0.812
##
##
## Covariance Coverage
## F2LOC1 F2LOC2 F2LOC3 F2LOC4
## ________ ________ ________ ________
## F2LOC1 0.820
## F2LOC2 0.816 0.819
## F2LOC3 0.815 0.814 0.817
## F2LOC4 0.812 0.811 0.813 0.814
##
##
## UNIVARIATE PROPORTIONS AND COUNTS FOR CATEGORICAL VARIABLES
##
## BYCONF1
## Category 1 0.009 106.372
## Category 2 0.066 789.544
## Category 3 0.580 6893.455
## Category 4 0.345 4099.427
## BYCONF2
## Category 1 0.013 147.092
## Category 2 0.067 788.727
## Category 3 0.517 6060.019
## Category 4 0.404 4735.196
## BYCONF3
## Category 1 0.009 100.960
## Category 2 0.072 844.180
## Category 3 0.525 6194.392
## Category 4 0.395 4654.544
## BYCONF4
## Category 1 0.017 199.173
## Category 2 0.106 1244.864
## Category 3 0.547 6448.019
## Category 4 0.331 3898.189
## BYLOC1
## Category 1 0.050 594.256
## Category 2 0.145 1723.431
## Category 3 0.483 5722.675
## Category 4 0.322 3815.344
## BYLOC2
## Category 1 0.026 309.484
## Category 2 0.087 1030.743
## Category 3 0.465 5495.124
## Category 4 0.422 4990.495
## BYLOC3
## Category 1 0.062 732.269
## Category 2 0.204 2415.727
## Category 3 0.566 6683.925
## Category 4 0.168 1983.910
## BYLOC4
## Category 1 0.055 653.835
## Category 2 0.135 1598.785
## Category 3 0.531 6286.881
## Category 4 0.279 3299.516
## F1CONF1
## Category 1 0.012 134.113
## Category 2 0.066 715.264
## Category 3 0.583 6319.218
## Category 4 0.338 3662.885
## F1CONF2
## Category 1 0.014 153.389
## Category 2 0.064 690.519
## Category 3 0.580 6259.005
## Category 4 0.342 3687.245
## F1CONF3
## Category 1 0.012 130.482
## Category 2 0.069 739.181
## Category 3 0.602 6465.546
## Category 4 0.317 3398.580
## F1CONF4
## Category 1 0.021 228.780
## Category 2 0.127 1371.274
## Category 3 0.578 6227.849
## Category 4 0.273 2941.364
## F1LOC1
## Category 1 0.043 458.400
## Category 2 0.175 1889.236
## Category 3 0.532 5734.112
## Category 4 0.250 2700.147
## F1LOC2
## Category 1 0.024 255.394
## Category 2 0.090 969.456
## Category 3 0.554 5977.981
## Category 4 0.332 3581.730
## F1LOC3
## Category 1 0.040 428.055
## Category 2 0.213 2297.721
## Category 3 0.595 6418.175
## Category 4 0.153 1646.682
## F1LOC4
## Category 1 0.036 386.089
## Category 2 0.169 1816.861
## Category 3 0.584 6269.453
## Category 4 0.211 2265.150
## F2CONF1
## Category 1 0.013 125.962
## Category 2 0.049 472.999
## Category 3 0.530 5100.972
## Category 4 0.408 3923.441
## F2CONF2
## Category 1 0.015 141.018
## Category 2 0.051 487.597
## Category 3 0.526 5016.276
## Category 4 0.408 3889.637
## F2CONF3
## Category 1 0.012 112.409
## Category 2 0.049 470.857
## Category 3 0.545 5193.745
## Category 4 0.394 3761.227
## F2CONF4
## Category 1 0.017 160.266
## Category 2 0.096 913.646
## Category 3 0.550 5253.880
## Category 4 0.337 3217.878
## F2LOC1
## Category 1 0.052 499.304
## Category 2 0.158 1515.287
## Category 3 0.518 4966.473
## Category 4 0.272 2601.878
## F2LOC2
## Category 1 0.024 230.374
## Category 2 0.076 729.473
## Category 3 0.556 5322.109
## Category 4 0.344 3291.022
## F2LOC3
## Category 1 0.038 362.924
## Category 2 0.185 1766.237
## Category 3 0.606 5796.676
## Category 4 0.171 1633.480
## F2LOC4
## Category 1 0.038 363.939
## Category 2 0.151 1441.183
## Category 3 0.578 5514.727
## Category 4 0.233 2218.804
##
##
## SAMPLE STATISTICS
##
##
## ESTIMATED SAMPLE STATISTICS
##
##
## MEANS/INTERCEPTS/THRESHOLDS
## BYCONF1$ BYCONF1$ BYCONF1$ BYCONF2$ BYCONF2$
## ________ ________ ________ ________ ________
## -2.368 -1.437 0.399 -2.240 -1.407
##
##
## MEANS/INTERCEPTS/THRESHOLDS
## BYCONF2$ BYCONF3$ BYCONF3$ BYCONF3$ BYCONF4$
## ________ ________ ________ ________ ________
## 0.244 -2.384 -1.404 0.267 -2.123
##
##
## MEANS/INTERCEPTS/THRESHOLDS
## BYCONF4$ BYCONF4$ BYLOC1$1 BYLOC1$2 BYLOC1$3
## ________ ________ ________ ________ ________
## -1.163 0.438 -1.644 -0.858 0.463
##
##
## MEANS/INTERCEPTS/THRESHOLDS
## BYLOC2$1 BYLOC2$2 BYLOC2$3 BYLOC3$1 BYLOC3$2
## ________ ________ ________ ________ ________
## -1.940 -1.209 0.197 -1.538 -0.624
##
##
## MEANS/INTERCEPTS/THRESHOLDS
## BYLOC3$3 BYLOC4$1 BYLOC4$2 BYLOC4$3 F1CONF1$
## ________ ________ ________ ________ ________
## 0.962 -1.596 -0.877 0.587 -2.245
##
##
## MEANS/INTERCEPTS/THRESHOLDS
## F1CONF1$ F1CONF1$ F1CONF2$ F1CONF2$ F1CONF2$
## ________ ________ ________ ________ ________
## -1.416 0.417 -2.191 -1.417 0.408
##
##
## MEANS/INTERCEPTS/THRESHOLDS
## F1CONF3$ F1CONF3$ F1CONF3$ F1CONF4$ F1CONF4$
## ________ ________ ________ ________ ________
## -2.252 -1.398 0.477 -2.029 -1.043
##
##
## MEANS/INTERCEPTS/THRESHOLDS
## F1CONF4$ F1LOC1$1 F1LOC1$2 F1LOC1$3 F1LOC2$1
## ________ ________ ________ ________ ________
## 0.603 -1.722 -0.780 0.673 -1.983
##
##
## MEANS/INTERCEPTS/THRESHOLDS
## F1LOC2$2 F1LOC2$3 F1LOC3$1 F1LOC3$2 F1LOC3$3
## ________ ________ ________ ________ ________
## -1.208 0.434 -1.755 -0.666 1.025
##
##
## MEANS/INTERCEPTS/THRESHOLDS
## F1LOC4$1 F1LOC4$2 F1LOC4$3 F2CONF1$ F2CONF1$
## ________ ________ ________ ________ ________
## -1.800 -0.823 0.803 -2.224 -1.536
##
##
## MEANS/INTERCEPTS/THRESHOLDS
## F2CONF1$ F2CONF2$ F2CONF2$ F2CONF2$ F2CONF3$
## ________ ________ ________ ________ ________
## 0.233 -2.176 -1.507 0.233 -2.264
##
##
## MEANS/INTERCEPTS/THRESHOLDS
## F2CONF3$ F2CONF3$ F2CONF4$ F2CONF4$ F2CONF4$
## ________ ________ ________ ________ ________
## -1.545 0.268 -2.125 -1.213 0.420
##
##
## MEANS/INTERCEPTS/THRESHOLDS
## F2LOC1$1 F2LOC1$2 F2LOC1$3 F2LOC2$1 F2LOC2$2
## ________ ________ ________ ________ ________
## -1.625 -0.806 0.608 -1.976 -1.280
##
##
## MEANS/INTERCEPTS/THRESHOLDS
## F2LOC2$3 F2LOC3$1 F2LOC3$2 F2LOC3$3 F2LOC4$1
## ________ ________ ________ ________ ________
## 0.402 -1.775 -0.763 0.951 -1.773
##
##
## MEANS/INTERCEPTS/THRESHOLDS
## F2LOC4$2 F2LOC4$3
## ________ ________
## -0.881 0.730
##
##
## CORRELATION MATRIX (WITH VARIANCES ON THE DIAGONAL)
## BYCONF1 BYCONF2 BYCONF3 BYCONF4 BYLOC1
## ________ ________ ________ ________ ________
## BYCONF1
## BYCONF2 0.467
## BYCONF3 0.389 0.516
## BYCONF4 0.661 0.500 0.425
## BYLOC1 0.274 0.220 0.190 0.314
## BYLOC2 0.090 0.208 0.142 0.180 0.328
## BYLOC3 0.277 0.230 0.226 0.336 0.371
## BYLOC4 0.322 0.288 0.261 0.397 0.381
## F1CONF1 0.475 0.292 0.257 0.358 0.165
## F1CONF2 0.317 0.368 0.234 0.298 0.199
## F1CONF3 0.270 0.266 0.345 0.282 0.188
## F1CONF4 0.352 0.261 0.202 0.375 0.186
## F1LOC1 0.216 0.136 0.136 0.218 0.294
## F1LOC2 0.047 0.115 0.085 0.100 0.194
## F1LOC3 0.198 0.186 0.167 0.232 0.240
## F1LOC4 0.227 0.208 0.187 0.284 0.245
## F2CONF1 0.429 0.210 0.239 0.321 0.147
## F2CONF2 0.266 0.268 0.182 0.245 0.152
## F2CONF3 0.264 0.242 0.298 0.246 0.119
## F2CONF4 0.325 0.201 0.179 0.325 0.201
## F2LOC1 0.137 0.097 0.078 0.177 0.242
## F2LOC2 0.043 0.091 0.055 0.073 0.183
## F2LOC3 0.186 0.172 0.138 0.191 0.236
## F2LOC4 0.183 0.149 0.081 0.208 0.215
##
##
## CORRELATION MATRIX (WITH VARIANCES ON THE DIAGONAL)
## BYLOC2 BYLOC3 BYLOC4 F1CONF1 F1CONF2
## ________ ________ ________ ________ ________
## BYLOC3 0.276
## BYLOC4 0.298 0.533
## F1CONF1 0.021 0.148 0.177
## F1CONF2 0.165 0.190 0.203 0.550
## F1CONF3 0.139 0.179 0.181 0.497 0.637
## F1CONF4 0.087 0.185 0.184 0.668 0.552
## F1LOC1 0.186 0.216 0.203 0.334 0.352
## F1LOC2 0.329 0.156 0.169 0.125 0.259
## F1LOC3 0.178 0.351 0.287 0.305 0.315
## F1LOC4 0.188 0.308 0.355 0.318 0.357
## F2CONF1 0.033 0.118 0.165 0.539 0.313
## F2CONF2 0.145 0.144 0.184 0.349 0.391
## F2CONF3 0.108 0.146 0.193 0.354 0.328
## F2CONF4 0.092 0.155 0.211 0.434 0.341
## F2LOC1 0.138 0.157 0.187 0.171 0.178
## F2LOC2 0.292 0.184 0.180 0.087 0.148
## F2LOC3 0.188 0.316 0.267 0.220 0.238
## F2LOC4 0.184 0.245 0.314 0.211 0.216
##
##
## CORRELATION MATRIX (WITH VARIANCES ON THE DIAGONAL)
## F1CONF3 F1CONF4 F1LOC1 F1LOC2 F1LOC3
## ________ ________ ________ ________ ________
## F1CONF4 0.512
## F1LOC1 0.301 0.366
## F1LOC2 0.203 0.173 0.421
## F1LOC3 0.278 0.384 0.444 0.324
## F1LOC4 0.341 0.415 0.467 0.376 0.606
## F2CONF1 0.290 0.435 0.245 0.062 0.230
## F2CONF2 0.294 0.299 0.250 0.177 0.198
## F2CONF3 0.434 0.316 0.207 0.161 0.190
## F2CONF4 0.321 0.446 0.309 0.169 0.280
## F2LOC1 0.157 0.204 0.305 0.205 0.221
## F2LOC2 0.137 0.173 0.214 0.375 0.198
## F2LOC3 0.244 0.279 0.287 0.222 0.426
## F2LOC4 0.197 0.249 0.261 0.241 0.362
##
##
## CORRELATION MATRIX (WITH VARIANCES ON THE DIAGONAL)
## F1LOC4 F2CONF1 F2CONF2 F2CONF3 F2CONF4
## ________ ________ ________ ________ ________
## F2CONF1 0.206
## F2CONF2 0.229 0.583
## F2CONF3 0.252 0.548 0.727
## F2CONF4 0.259 0.702 0.587 0.549
## F2LOC1 0.261 0.278 0.266 0.248 0.332
## F2LOC2 0.247 0.120 0.235 0.204 0.205
## F2LOC3 0.328 0.291 0.297 0.254 0.393
## F2LOC4 0.399 0.297 0.342 0.306 0.418
##
##
## CORRELATION MATRIX (WITH VARIANCES ON THE DIAGONAL)
## F2LOC1 F2LOC2 F2LOC3 F2LOC4
## ________ ________ ________ ________
## F2LOC2 0.453
## F2LOC3 0.468 0.405
## F2LOC4 0.486 0.442 0.655
##
##
## THE MODEL ESTIMATION TERMINATED NORMALLY
##
##
##
## MODEL FIT INFORMATION
##
## Number of Free Parameters 111
##
## Chi-Square Test of Model Fit
##
## Value 1771.108*
## Degrees of Freedom 237
## P-Value 0.0000
##
## * The chi-square value for MLM, MLMV, MLR, ULSMV, WLSM and WLSMV cannot be used
## for chi-square difference testing in the regular way. MLM, MLR and WLSM
## chi-square difference testing is described on the Mplus website. MLMV, WLSMV,
## and ULSMV difference testing is done using the DIFFTEST option.
##
## RMSEA (Root Mean Square Error Of Approximation)
##
## Estimate 0.023
## 90 Percent C.I. 0.022 0.024
## Probability RMSEA <= .05 1.000
##
## CFI/TLI
##
## CFI 0.964
## TLI 0.958
##
## Chi-Square Test of Model Fit for the Baseline Model
##
## Value 42593.921
## Degrees of Freedom 276
## P-Value 0.0000
##
## SRMR (Standardized Root Mean Square Residual)
##
## Value 0.042
##
## Optimum Function Value for Weighted Least-Squares Estimator
##
## Value 0.10585169D+00
##
##
##
## MODEL RESULTS
##
## Two-Tailed
## Estimate S.E. Est./S.E. P-Value
##
## F1 BY
## BYCONF1 1.000 0.000 999.000 999.000
## BYCONF2 0.850 0.018 47.304 0.000
## BYCONF3 0.763 0.020 37.637 0.000
## BYCONF4 1.032 0.018 55.868 0.000
##
## F2 BY
## BYLOC1 1.000 0.000 999.000 999.000
## BYLOC2 0.757 0.034 22.422 0.000
## BYLOC3 1.120 0.041 27.613 0.000
## BYLOC4 1.205 0.041 29.420 0.000
##
## F3 BY
## F1CONF1 1.000 0.000 999.000 999.000
## F1CONF2 0.964 0.016 59.388 0.000
## F1CONF3 0.906 0.017 52.304 0.000
## F1CONF4 1.007 0.017 58.364 0.000
##
## F4 BY
## F1LOC1 1.000 0.000 999.000 999.000
## F1LOC2 0.744 0.028 26.879 0.000
## F1LOC3 1.099 0.024 45.175 0.000
## F1LOC4 1.165 0.030 38.456 0.000
##
## F5 BY
## F2CONF1 1.000 0.000 999.000 999.000
## F2CONF2 1.022 0.021 49.680 0.000
## F2CONF3 0.993 0.017 59.446 0.000
## F2CONF4 1.028 0.017 60.655 0.000
##
## F6 BY
## F2LOC1 1.000 0.000 999.000 999.000
## F2LOC2 0.878 0.031 28.507 0.000
## F2LOC3 1.227 0.039 31.809 0.000
## F2LOC4 1.267 0.036 35.336 0.000
##
## F2 WITH
## F1 0.273 0.011 25.286 0.000
##
## F3 WITH
## F1 0.358 0.012 30.455 0.000
## F2 0.165 0.011 15.535 0.000
##
## F4 WITH
## F1 0.195 0.011 18.005 0.000
## F2 0.234 0.011 22.069 0.000
## F3 0.316 0.011 27.821 0.000
##
## F5 WITH
## F1 0.291 0.011 25.616 0.000
## F2 0.143 0.011 13.501 0.000
## F3 0.385 0.012 32.554 0.000
## F4 0.213 0.011 18.621 0.000
##
## F6 WITH
## F1 0.133 0.011 11.823 0.000
## F2 0.196 0.012 16.116 0.000
## F3 0.185 0.012 15.288 0.000
## F4 0.258 0.011 24.290 0.000
## F5 0.258 0.010 25.873 0.000
##
## Thresholds
## BYCONF1$1 -2.368 0.053 -44.587 0.000
## BYCONF1$2 -1.437 0.026 -55.700 0.000
## BYCONF1$3 0.399 0.020 20.135 0.000
## BYCONF2$1 -2.240 0.043 -51.587 0.000
## BYCONF2$2 -1.407 0.026 -54.699 0.000
## BYCONF2$3 0.244 0.019 12.587 0.000
## BYCONF3$1 -2.384 0.046 -52.129 0.000
## BYCONF3$2 -1.404 0.027 -51.756 0.000
## BYCONF3$3 0.267 0.020 13.490 0.000
## BYCONF4$1 -2.123 0.039 -54.659 0.000
## BYCONF4$2 -1.163 0.026 -45.302 0.000
## BYCONF4$3 0.438 0.020 22.231 0.000
## BYLOC1$1 -1.644 0.039 -42.179 0.000
## BYLOC1$2 -0.858 0.023 -37.460 0.000
## BYLOC1$3 0.463 0.019 23.795 0.000
## BYLOC2$1 -1.940 0.034 -56.444 0.000
## BYLOC2$2 -1.209 0.028 -42.872 0.000
## BYLOC2$3 0.197 0.019 10.203 0.000
## BYLOC3$1 -1.538 0.030 -51.782 0.000
## BYLOC3$2 -0.624 0.020 -31.493 0.000
## BYLOC3$3 0.962 0.026 37.009 0.000
## BYLOC4$1 -1.596 0.032 -49.727 0.000
## BYLOC4$2 -0.877 0.022 -40.654 0.000
## BYLOC4$3 0.587 0.020 28.988 0.000
## F1CONF1$1 -2.245 0.046 -49.252 0.000
## F1CONF1$2 -1.416 0.025 -55.701 0.000
## F1CONF1$3 0.417 0.020 21.034 0.000
## F1CONF2$1 -2.191 0.061 -36.218 0.000
## F1CONF2$2 -1.417 0.027 -52.658 0.000
## F1CONF2$3 0.408 0.019 21.012 0.000
## F1CONF3$1 -2.252 0.072 -31.068 0.000
## F1CONF3$2 -1.398 0.032 -44.300 0.000
## F1CONF3$3 0.477 0.019 24.712 0.000
## F1CONF4$1 -2.029 0.052 -39.351 0.000
## F1CONF4$2 -1.043 0.023 -44.429 0.000
## F1CONF4$3 0.603 0.020 29.584 0.000
## F1LOC1$1 -1.722 0.042 -40.941 0.000
## F1LOC1$2 -0.780 0.021 -36.949 0.000
## F1LOC1$3 0.673 0.020 34.413 0.000
## F1LOC2$1 -1.983 0.056 -35.311 0.000
## F1LOC2$2 -1.208 0.026 -47.097 0.000
## F1LOC2$3 0.434 0.020 22.098 0.000
## F1LOC3$1 -1.755 0.041 -43.214 0.000
## F1LOC3$2 -0.666 0.021 -32.446 0.000
## F1LOC3$3 1.025 0.024 42.007 0.000
## F1LOC4$1 -1.800 0.041 -44.037 0.000
## F1LOC4$2 -0.823 0.021 -38.294 0.000
## F1LOC4$3 0.803 0.022 36.513 0.000
## F2CONF1$1 -2.224 0.072 -30.695 0.000
## F2CONF1$2 -1.536 0.029 -53.809 0.000
## F2CONF1$3 0.233 0.021 11.253 0.000
## F2CONF2$1 -2.176 0.043 -50.910 0.000
## F2CONF2$2 -1.507 0.030 -50.859 0.000
## F2CONF2$3 0.233 0.020 11.414 0.000
## F2CONF3$1 -2.264 0.074 -30.752 0.000
## F2CONF3$2 -1.545 0.031 -49.589 0.000
## F2CONF3$3 0.268 0.021 13.010 0.000
## F2CONF4$1 -2.125 0.058 -36.473 0.000
## F2CONF4$2 -1.213 0.029 -41.846 0.000
## F2CONF4$3 0.420 0.020 20.741 0.000
## F2LOC1$1 -1.625 0.040 -40.446 0.000
## F2LOC1$2 -0.806 0.025 -32.859 0.000
## F2LOC1$3 0.608 0.021 29.253 0.000
## F2LOC2$1 -1.976 0.036 -54.645 0.000
## F2LOC2$2 -1.280 0.030 -42.353 0.000
## F2LOC2$3 0.402 0.022 18.630 0.000
## F2LOC3$1 -1.775 0.040 -44.519 0.000
## F2LOC3$2 -0.763 0.024 -31.622 0.000
## F2LOC3$3 0.951 0.024 39.988 0.000
## F2LOC4$1 -1.773 0.045 -39.047 0.000
## F2LOC4$2 -0.881 0.027 -32.113 0.000
## F2LOC4$3 0.730 0.023 31.455 0.000
##
## Variances
## F1 0.611 0.015 39.609 0.000
## F2 0.359 0.020 18.108 0.000
## F3 0.622 0.014 43.120 0.000
## F4 0.440 0.018 24.821 0.000
## F5 0.624 0.016 39.777 0.000
## F6 0.410 0.022 18.588 0.000
##
##
## STANDARDIZED MODEL RESULTS
##
##
## STDYX Standardization
##
## Two-Tailed
## Estimate S.E. Est./S.E. P-Value
##
## F1 BY
## BYCONF1 0.782 0.010 79.218 0.000
## BYCONF2 0.664 0.011 58.280 0.000
## BYCONF3 0.597 0.014 42.722 0.000
## BYCONF4 0.807 0.009 92.360 0.000
##
## F2 BY
## BYLOC1 0.599 0.017 36.215 0.000
## BYLOC2 0.453 0.016 27.774 0.000
## BYLOC3 0.671 0.013 52.277 0.000
## BYLOC4 0.722 0.012 57.839 0.000
##
## F3 BY
## F1CONF1 0.789 0.009 86.240 0.000
## F1CONF2 0.760 0.010 72.685 0.000
## F1CONF3 0.715 0.011 63.428 0.000
## F1CONF4 0.795 0.010 80.472 0.000
##
## F4 BY
## F1LOC1 0.663 0.013 49.642 0.000
## F1LOC2 0.494 0.015 32.206 0.000
## F1LOC3 0.729 0.011 63.446 0.000
## F1LOC4 0.773 0.011 68.445 0.000
##
## F5 BY
## F2CONF1 0.790 0.010 79.555 0.000
## F2CONF2 0.808 0.011 72.967 0.000
## F2CONF3 0.785 0.010 81.201 0.000
## F2CONF4 0.813 0.010 82.522 0.000
##
## F6 BY
## F2LOC1 0.640 0.017 37.175 0.000
## F2LOC2 0.562 0.015 38.326 0.000
## F2LOC3 0.786 0.011 73.188 0.000
## F2LOC4 0.811 0.013 60.726 0.000
##
## F2 WITH
## F1 0.583 0.015 40.143 0.000
##
## F3 WITH
## F1 0.581 0.016 36.686 0.000
## F2 0.349 0.020 17.559 0.000
##
## F4 WITH
## F1 0.375 0.018 21.339 0.000
## F2 0.589 0.019 31.446 0.000
## F3 0.603 0.015 39.265 0.000
##
## F5 WITH
## F1 0.470 0.016 30.124 0.000
## F2 0.301 0.021 14.602 0.000
## F3 0.618 0.014 45.656 0.000
## F4 0.407 0.017 23.701 0.000
##
## F6 WITH
## F1 0.266 0.020 13.259 0.000
## F2 0.511 0.021 24.072 0.000
## F3 0.366 0.020 18.450 0.000
## F4 0.608 0.015 40.031 0.000
## F5 0.510 0.014 37.162 0.000
##
## Thresholds
## BYCONF1$1 -2.368 0.053 -44.587 0.000
## BYCONF1$2 -1.437 0.026 -55.700 0.000
## BYCONF1$3 0.399 0.020 20.135 0.000
## BYCONF2$1 -2.240 0.043 -51.587 0.000
## BYCONF2$2 -1.407 0.026 -54.699 0.000
## BYCONF2$3 0.244 0.019 12.587 0.000
## BYCONF3$1 -2.384 0.046 -52.129 0.000
## BYCONF3$2 -1.404 0.027 -51.756 0.000
## BYCONF3$3 0.267 0.020 13.490 0.000
## BYCONF4$1 -2.123 0.039 -54.659 0.000
## BYCONF4$2 -1.163 0.026 -45.302 0.000
## BYCONF4$3 0.438 0.020 22.231 0.000
## BYLOC1$1 -1.644 0.039 -42.179 0.000
## BYLOC1$2 -0.858 0.023 -37.460 0.000
## BYLOC1$3 0.463 0.019 23.795 0.000
## BYLOC2$1 -1.940 0.034 -56.444 0.000
## BYLOC2$2 -1.209 0.028 -42.872 0.000
## BYLOC2$3 0.197 0.019 10.203 0.000
## BYLOC3$1 -1.538 0.030 -51.782 0.000
## BYLOC3$2 -0.624 0.020 -31.493 0.000
## BYLOC3$3 0.962 0.026 37.009 0.000
## BYLOC4$1 -1.596 0.032 -49.727 0.000
## BYLOC4$2 -0.877 0.022 -40.654 0.000
## BYLOC4$3 0.587 0.020 28.988 0.000
## F1CONF1$1 -2.245 0.046 -49.252 0.000
## F1CONF1$2 -1.416 0.025 -55.701 0.000
## F1CONF1$3 0.417 0.020 21.034 0.000
## F1CONF2$1 -2.191 0.061 -36.218 0.000
## F1CONF2$2 -1.417 0.027 -52.658 0.000
## F1CONF2$3 0.408 0.019 21.012 0.000
## F1CONF3$1 -2.252 0.072 -31.068 0.000
## F1CONF3$2 -1.398 0.032 -44.300 0.000
## F1CONF3$3 0.477 0.019 24.712 0.000
## F1CONF4$1 -2.029 0.052 -39.351 0.000
## F1CONF4$2 -1.043 0.023 -44.429 0.000
## F1CONF4$3 0.603 0.020 29.584 0.000
## F1LOC1$1 -1.722 0.042 -40.941 0.000
## F1LOC1$2 -0.780 0.021 -36.949 0.000
## F1LOC1$3 0.673 0.020 34.413 0.000
## F1LOC2$1 -1.983 0.056 -35.311 0.000
## F1LOC2$2 -1.208 0.026 -47.097 0.000
## F1LOC2$3 0.434 0.020 22.098 0.000
## F1LOC3$1 -1.755 0.041 -43.214 0.000
## F1LOC3$2 -0.666 0.021 -32.446 0.000
## F1LOC3$3 1.025 0.024 42.007 0.000
## F1LOC4$1 -1.800 0.041 -44.037 0.000
## F1LOC4$2 -0.823 0.021 -38.294 0.000
## F1LOC4$3 0.803 0.022 36.513 0.000
## F2CONF1$1 -2.224 0.072 -30.695 0.000
## F2CONF1$2 -1.536 0.029 -53.809 0.000
## F2CONF1$3 0.233 0.021 11.253 0.000
## F2CONF2$1 -2.176 0.043 -50.910 0.000
## F2CONF2$2 -1.507 0.030 -50.859 0.000
## F2CONF2$3 0.233 0.020 11.414 0.000
## F2CONF3$1 -2.264 0.074 -30.752 0.000
## F2CONF3$2 -1.545 0.031 -49.589 0.000
## F2CONF3$3 0.268 0.021 13.010 0.000
## F2CONF4$1 -2.125 0.058 -36.473 0.000
## F2CONF4$2 -1.213 0.029 -41.846 0.000
## F2CONF4$3 0.420 0.020 20.741 0.000
## F2LOC1$1 -1.625 0.040 -40.446 0.000
## F2LOC1$2 -0.806 0.025 -32.859 0.000
## F2LOC1$3 0.608 0.021 29.253 0.000
## F2LOC2$1 -1.976 0.036 -54.645 0.000
## F2LOC2$2 -1.280 0.030 -42.353 0.000
## F2LOC2$3 0.402 0.022 18.630 0.000
## F2LOC3$1 -1.775 0.040 -44.519 0.000
## F2LOC3$2 -0.763 0.024 -31.622 0.000
## F2LOC3$3 0.951 0.024 39.988 0.000
## F2LOC4$1 -1.773 0.045 -39.047 0.000
## F2LOC4$2 -0.881 0.027 -32.113 0.000
## F2LOC4$3 0.730 0.023 31.455 0.000
##
## Variances
## F1 1.000 0.000 999.000 999.000
## F2 1.000 0.000 999.000 999.000
## F3 1.000 0.000 999.000 999.000
## F4 1.000 0.000 999.000 999.000
## F5 1.000 0.000 999.000 999.000
## F6 1.000 0.000 999.000 999.000
##
##
## STDY Standardization
##
## Two-Tailed
## Estimate S.E. Est./S.E. P-Value
##
## F1 BY
## BYCONF1 0.782 0.010 79.218 0.000
## BYCONF2 0.664 0.011 58.280 0.000
## BYCONF3 0.597 0.014 42.722 0.000
## BYCONF4 0.807 0.009 92.360 0.000
##
## F2 BY
## BYLOC1 0.599 0.017 36.215 0.000
## BYLOC2 0.453 0.016 27.774 0.000
## BYLOC3 0.671 0.013 52.277 0.000
## BYLOC4 0.722 0.012 57.839 0.000
##
## F3 BY
## F1CONF1 0.789 0.009 86.240 0.000
## F1CONF2 0.760 0.010 72.685 0.000
## F1CONF3 0.715 0.011 63.428 0.000
## F1CONF4 0.795 0.010 80.472 0.000
##
## F4 BY
## F1LOC1 0.663 0.013 49.642 0.000
## F1LOC2 0.494 0.015 32.206 0.000
## F1LOC3 0.729 0.011 63.446 0.000
## F1LOC4 0.773 0.011 68.445 0.000
##
## F5 BY
## F2CONF1 0.790 0.010 79.555 0.000
## F2CONF2 0.808 0.011 72.967 0.000
## F2CONF3 0.785 0.010 81.201 0.000
## F2CONF4 0.813 0.010 82.522 0.000
##
## F6 BY
## F2LOC1 0.640 0.017 37.175 0.000
## F2LOC2 0.562 0.015 38.326 0.000
## F2LOC3 0.786 0.011 73.188 0.000
## F2LOC4 0.811 0.013 60.726 0.000
##
## F2 WITH
## F1 0.583 0.015 40.143 0.000
##
## F3 WITH
## F1 0.581 0.016 36.686 0.000
## F2 0.349 0.020 17.559 0.000
##
## F4 WITH
## F1 0.375 0.018 21.339 0.000
## F2 0.589 0.019 31.446 0.000
## F3 0.603 0.015 39.265 0.000
##
## F5 WITH
## F1 0.470 0.016 30.124 0.000
## F2 0.301 0.021 14.602 0.000
## F3 0.618 0.014 45.656 0.000
## F4 0.407 0.017 23.701 0.000
##
## F6 WITH
## F1 0.266 0.020 13.259 0.000
## F2 0.511 0.021 24.072 0.000
## F3 0.366 0.020 18.450 0.000
## F4 0.608 0.015 40.031 0.000
## F5 0.510 0.014 37.162 0.000
##
## Thresholds
## BYCONF1$1 -2.368 0.053 -44.587 0.000
## BYCONF1$2 -1.437 0.026 -55.700 0.000
## BYCONF1$3 0.399 0.020 20.135 0.000
## BYCONF2$1 -2.240 0.043 -51.587 0.000
## BYCONF2$2 -1.407 0.026 -54.699 0.000
## BYCONF2$3 0.244 0.019 12.587 0.000
## BYCONF3$1 -2.384 0.046 -52.129 0.000
## BYCONF3$2 -1.404 0.027 -51.756 0.000
## BYCONF3$3 0.267 0.020 13.490 0.000
## BYCONF4$1 -2.123 0.039 -54.659 0.000
## BYCONF4$2 -1.163 0.026 -45.302 0.000
## BYCONF4$3 0.438 0.020 22.231 0.000
## BYLOC1$1 -1.644 0.039 -42.179 0.000
## BYLOC1$2 -0.858 0.023 -37.460 0.000
## BYLOC1$3 0.463 0.019 23.795 0.000
## BYLOC2$1 -1.940 0.034 -56.444 0.000
## BYLOC2$2 -1.209 0.028 -42.872 0.000
## BYLOC2$3 0.197 0.019 10.203 0.000
## BYLOC3$1 -1.538 0.030 -51.782 0.000
## BYLOC3$2 -0.624 0.020 -31.493 0.000
## BYLOC3$3 0.962 0.026 37.009 0.000
## BYLOC4$1 -1.596 0.032 -49.727 0.000
## BYLOC4$2 -0.877 0.022 -40.654 0.000
## BYLOC4$3 0.587 0.020 28.988 0.000
## F1CONF1$1 -2.245 0.046 -49.252 0.000
## F1CONF1$2 -1.416 0.025 -55.701 0.000
## F1CONF1$3 0.417 0.020 21.034 0.000
## F1CONF2$1 -2.191 0.061 -36.218 0.000
## F1CONF2$2 -1.417 0.027 -52.658 0.000
## F1CONF2$3 0.408 0.019 21.012 0.000
## F1CONF3$1 -2.252 0.072 -31.068 0.000
## F1CONF3$2 -1.398 0.032 -44.300 0.000
## F1CONF3$3 0.477 0.019 24.712 0.000
## F1CONF4$1 -2.029 0.052 -39.351 0.000
## F1CONF4$2 -1.043 0.023 -44.429 0.000
## F1CONF4$3 0.603 0.020 29.584 0.000
## F1LOC1$1 -1.722 0.042 -40.941 0.000
## F1LOC1$2 -0.780 0.021 -36.949 0.000
## F1LOC1$3 0.673 0.020 34.413 0.000
## F1LOC2$1 -1.983 0.056 -35.311 0.000
## F1LOC2$2 -1.208 0.026 -47.097 0.000
## F1LOC2$3 0.434 0.020 22.098 0.000
## F1LOC3$1 -1.755 0.041 -43.214 0.000
## F1LOC3$2 -0.666 0.021 -32.446 0.000
## F1LOC3$3 1.025 0.024 42.007 0.000
## F1LOC4$1 -1.800 0.041 -44.037 0.000
## F1LOC4$2 -0.823 0.021 -38.294 0.000
## F1LOC4$3 0.803 0.022 36.513 0.000
## F2CONF1$1 -2.224 0.072 -30.695 0.000
## F2CONF1$2 -1.536 0.029 -53.809 0.000
## F2CONF1$3 0.233 0.021 11.253 0.000
## F2CONF2$1 -2.176 0.043 -50.910 0.000
## F2CONF2$2 -1.507 0.030 -50.859 0.000
## F2CONF2$3 0.233 0.020 11.414 0.000
## F2CONF3$1 -2.264 0.074 -30.752 0.000
## F2CONF3$2 -1.545 0.031 -49.589 0.000
## F2CONF3$3 0.268 0.021 13.010 0.000
## F2CONF4$1 -2.125 0.058 -36.473 0.000
## F2CONF4$2 -1.213 0.029 -41.846 0.000
## F2CONF4$3 0.420 0.020 20.741 0.000
## F2LOC1$1 -1.625 0.040 -40.446 0.000
## F2LOC1$2 -0.806 0.025 -32.859 0.000
## F2LOC1$3 0.608 0.021 29.253 0.000
## F2LOC2$1 -1.976 0.036 -54.645 0.000
## F2LOC2$2 -1.280 0.030 -42.353 0.000
## F2LOC2$3 0.402 0.022 18.630 0.000
## F2LOC3$1 -1.775 0.040 -44.519 0.000
## F2LOC3$2 -0.763 0.024 -31.622 0.000
## F2LOC3$3 0.951 0.024 39.988 0.000
## F2LOC4$1 -1.773 0.045 -39.047 0.000
## F2LOC4$2 -0.881 0.027 -32.113 0.000
## F2LOC4$3 0.730 0.023 31.455 0.000
##
## Variances
## F1 1.000 0.000 999.000 999.000
## F2 1.000 0.000 999.000 999.000
## F3 1.000 0.000 999.000 999.000
## F4 1.000 0.000 999.000 999.000
## F5 1.000 0.000 999.000 999.000
## F6 1.000 0.000 999.000 999.000
##
##
## STD Standardization
##
## Two-Tailed
## Estimate S.E. Est./S.E. P-Value
##
## F1 BY
## BYCONF1 0.782 0.010 79.218 0.000
## BYCONF2 0.664 0.011 58.280 0.000
## BYCONF3 0.597 0.014 42.722 0.000
## BYCONF4 0.807 0.009 92.360 0.000
##
## F2 BY
## BYLOC1 0.599 0.017 36.215 0.000
## BYLOC2 0.453 0.016 27.774 0.000
## BYLOC3 0.671 0.013 52.277 0.000
## BYLOC4 0.722 0.012 57.839 0.000
##
## F3 BY
## F1CONF1 0.789 0.009 86.240 0.000
## F1CONF2 0.760 0.010 72.685 0.000
## F1CONF3 0.715 0.011 63.428 0.000
## F1CONF4 0.795 0.010 80.472 0.000
##
## F4 BY
## F1LOC1 0.663 0.013 49.642 0.000
## F1LOC2 0.494 0.015 32.206 0.000
## F1LOC3 0.729 0.011 63.446 0.000
## F1LOC4 0.773 0.011 68.445 0.000
##
## F5 BY
## F2CONF1 0.790 0.010 79.555 0.000
## F2CONF2 0.808 0.011 72.967 0.000
## F2CONF3 0.785 0.010 81.201 0.000
## F2CONF4 0.813 0.010 82.522 0.000
##
## F6 BY
## F2LOC1 0.640 0.017 37.175 0.000
## F2LOC2 0.562 0.015 38.326 0.000
## F2LOC3 0.786 0.011 73.188 0.000
## F2LOC4 0.811 0.013 60.726 0.000
##
## F2 WITH
## F1 0.583 0.015 40.143 0.000
##
## F3 WITH
## F1 0.581 0.016 36.686 0.000
## F2 0.349 0.020 17.559 0.000
##
## F4 WITH
## F1 0.375 0.018 21.339 0.000
## F2 0.589 0.019 31.446 0.000
## F3 0.603 0.015 39.265 0.000
##
## F5 WITH
## F1 0.470 0.016 30.124 0.000
## F2 0.301 0.021 14.602 0.000
## F3 0.618 0.014 45.656 0.000
## F4 0.407 0.017 23.701 0.000
##
## F6 WITH
## F1 0.266 0.020 13.259 0.000
## F2 0.511 0.021 24.072 0.000
## F3 0.366 0.020 18.450 0.000
## F4 0.608 0.015 40.031 0.000
## F5 0.510 0.014 37.162 0.000
##
## Thresholds
## BYCONF1$1 -2.368 0.053 -44.587 0.000
## BYCONF1$2 -1.437 0.026 -55.700 0.000
## BYCONF1$3 0.399 0.020 20.135 0.000
## BYCONF2$1 -2.240 0.043 -51.587 0.000
## BYCONF2$2 -1.407 0.026 -54.699 0.000
## BYCONF2$3 0.244 0.019 12.587 0.000
## BYCONF3$1 -2.384 0.046 -52.129 0.000
## BYCONF3$2 -1.404 0.027 -51.756 0.000
## BYCONF3$3 0.267 0.020 13.490 0.000
## BYCONF4$1 -2.123 0.039 -54.659 0.000
## BYCONF4$2 -1.163 0.026 -45.302 0.000
## BYCONF4$3 0.438 0.020 22.231 0.000
## BYLOC1$1 -1.644 0.039 -42.179 0.000
## BYLOC1$2 -0.858 0.023 -37.460 0.000
## BYLOC1$3 0.463 0.019 23.795 0.000
## BYLOC2$1 -1.940 0.034 -56.444 0.000
## BYLOC2$2 -1.209 0.028 -42.872 0.000
## BYLOC2$3 0.197 0.019 10.203 0.000
## BYLOC3$1 -1.538 0.030 -51.782 0.000
## BYLOC3$2 -0.624 0.020 -31.493 0.000
## BYLOC3$3 0.962 0.026 37.009 0.000
## BYLOC4$1 -1.596 0.032 -49.727 0.000
## BYLOC4$2 -0.877 0.022 -40.654 0.000
## BYLOC4$3 0.587 0.020 28.988 0.000
## F1CONF1$1 -2.245 0.046 -49.252 0.000
## F1CONF1$2 -1.416 0.025 -55.701 0.000
## F1CONF1$3 0.417 0.020 21.034 0.000
## F1CONF2$1 -2.191 0.061 -36.218 0.000
## F1CONF2$2 -1.417 0.027 -52.658 0.000
## F1CONF2$3 0.408 0.019 21.012 0.000
## F1CONF3$1 -2.252 0.072 -31.068 0.000
## F1CONF3$2 -1.398 0.032 -44.300 0.000
## F1CONF3$3 0.477 0.019 24.712 0.000
## F1CONF4$1 -2.029 0.052 -39.351 0.000
## F1CONF4$2 -1.043 0.023 -44.429 0.000
## F1CONF4$3 0.603 0.020 29.584 0.000
## F1LOC1$1 -1.722 0.042 -40.941 0.000
## F1LOC1$2 -0.780 0.021 -36.949 0.000
## F1LOC1$3 0.673 0.020 34.413 0.000
## F1LOC2$1 -1.983 0.056 -35.311 0.000
## F1LOC2$2 -1.208 0.026 -47.097 0.000
## F1LOC2$3 0.434 0.020 22.098 0.000
## F1LOC3$1 -1.755 0.041 -43.214 0.000
## F1LOC3$2 -0.666 0.021 -32.446 0.000
## F1LOC3$3 1.025 0.024 42.007 0.000
## F1LOC4$1 -1.800 0.041 -44.037 0.000
## F1LOC4$2 -0.823 0.021 -38.294 0.000
## F1LOC4$3 0.803 0.022 36.513 0.000
## F2CONF1$1 -2.224 0.072 -30.695 0.000
## F2CONF1$2 -1.536 0.029 -53.809 0.000
## F2CONF1$3 0.233 0.021 11.253 0.000
## F2CONF2$1 -2.176 0.043 -50.910 0.000
## F2CONF2$2 -1.507 0.030 -50.859 0.000
## F2CONF2$3 0.233 0.020 11.414 0.000
## F2CONF3$1 -2.264 0.074 -30.752 0.000
## F2CONF3$2 -1.545 0.031 -49.589 0.000
## F2CONF3$3 0.268 0.021 13.010 0.000
## F2CONF4$1 -2.125 0.058 -36.473 0.000
## F2CONF4$2 -1.213 0.029 -41.846 0.000
## F2CONF4$3 0.420 0.020 20.741 0.000
## F2LOC1$1 -1.625 0.040 -40.446 0.000
## F2LOC1$2 -0.806 0.025 -32.859 0.000
## F2LOC1$3 0.608 0.021 29.253 0.000
## F2LOC2$1 -1.976 0.036 -54.645 0.000
## F2LOC2$2 -1.280 0.030 -42.353 0.000
## F2LOC2$3 0.402 0.022 18.630 0.000
## F2LOC3$1 -1.775 0.040 -44.519 0.000
## F2LOC3$2 -0.763 0.024 -31.622 0.000
## F2LOC3$3 0.951 0.024 39.988 0.000
## F2LOC4$1 -1.773 0.045 -39.047 0.000
## F2LOC4$2 -0.881 0.027 -32.113 0.000
## F2LOC4$3 0.730 0.023 31.455 0.000
##
## Variances
## F1 1.000 0.000 999.000 999.000
## F2 1.000 0.000 999.000 999.000
## F3 1.000 0.000 999.000 999.000
## F4 1.000 0.000 999.000 999.000
## F5 1.000 0.000 999.000 999.000
## F6 1.000 0.000 999.000 999.000
##
##
## R-SQUARE
##
## Observed Two-Tailed Residual
## Variable Estimate S.E. Est./S.E. P-Value Variance
##
## BYCONF1 0.611 0.015 39.609 0.000 0.389
## BYCONF2 0.441 0.015 29.140 0.000 0.559
## BYCONF3 0.356 0.017 21.361 0.000 0.644
## BYCONF4 0.651 0.014 46.180 0.000 0.349
## BYLOC1 0.359 0.020 18.108 0.000 0.641
## BYLOC2 0.206 0.015 13.887 0.000 0.794
## BYLOC3 0.450 0.017 26.138 0.000 0.550
## BYLOC4 0.521 0.018 28.920 0.000 0.479
## F1CONF1 0.622 0.014 43.120 0.000 0.378
## F1CONF2 0.578 0.016 36.343 0.000 0.422
## F1CONF3 0.511 0.016 31.714 0.000 0.489
## F1CONF4 0.631 0.016 40.236 0.000 0.369
## F1LOC1 0.440 0.018 24.821 0.000 0.560
## F1LOC2 0.244 0.015 16.103 0.000 0.756
## F1LOC3 0.531 0.017 31.723 0.000 0.469
## F1LOC4 0.598 0.017 34.222 0.000 0.402
## F2CONF1 0.624 0.016 39.777 0.000 0.376
## F2CONF2 0.652 0.018 36.483 0.000 0.348
## F2CONF3 0.616 0.015 40.601 0.000 0.384
## F2CONF4 0.660 0.016 41.261 0.000 0.340
## F2LOC1 0.410 0.022 18.588 0.000 0.590
## F2LOC2 0.316 0.016 19.163 0.000 0.684
## F2LOC3 0.617 0.017 36.594 0.000 0.383
## F2LOC4 0.658 0.022 30.363 0.000 0.342
##
##
## QUALITY OF NUMERICAL RESULTS
##
## Condition Number for the Information Matrix 0.281E-02
## (ratio of smallest to largest eigenvalue)
##
##
## MODEL MODIFICATION INDICES
##
## NOTE: Modification indices for direct effects of observed dependent variables
## regressed on covariates and residual covariances among observed dependent
## variables may not be included. To include these, request MODINDICES (ALL).
##
## Minimum M.I. value for printing the modification index 10.000
##
## M.I. E.P.C. Std E.P.C. StdYX E.P.C.
##
## BY Statements
##
## F1 BY BYLOC2 22.309 -0.148 -0.116 -0.116
## F1 BY BYLOC4 10.201 0.110 0.086 0.086
## F1 BY F1CONF1 10.261 0.107 0.083 0.083
## F1 BY F1LOC2 30.786 -0.145 -0.113 -0.113
## F1 BY F2CONF1 22.668 0.141 0.110 0.110
## F1 BY F2CONF2 21.086 -0.133 -0.104 -0.104
## F1 BY F2LOC2 18.687 -0.102 -0.080 -0.080
## F2 BY BYCONF1 14.133 -0.168 -0.101 -0.101
## F2 BY BYCONF4 24.305 0.223 0.134 0.134
## F2 BY F1CONF1 14.933 -0.141 -0.084 -0.084
## F2 BY F2CONF4 32.015 0.194 0.116 0.116
## F3 BY BYCONF1 15.419 0.127 0.100 0.100
## F3 BY BYLOC1 14.011 0.091 0.072 0.072
## F3 BY F1LOC1 35.132 0.181 0.143 0.143
## F3 BY F1LOC2 41.420 -0.188 -0.148 -0.148
## F3 BY F2CONF1 30.542 0.197 0.155 0.155
## F3 BY F2CONF2 37.514 -0.213 -0.168 -0.168
## F3 BY F2LOC2 16.626 -0.096 -0.076 -0.076
## F4 BY BYLOC4 12.213 -0.161 -0.107 -0.107
## F4 BY F1CONF1 48.241 -0.266 -0.176 -0.176
## F4 BY F1CONF4 14.791 0.146 0.097 0.097
## F4 BY F2CONF3 11.576 -0.110 -0.073 -0.073
## F4 BY F2CONF4 53.370 0.235 0.156 0.156
## F5 BY BYCONF1 17.419 0.119 0.094 0.094
## F5 BY F1CONF1 18.743 0.159 0.125 0.125
## F5 BY F1LOC1 27.882 0.142 0.112 0.112
## F5 BY F1LOC2 11.048 -0.088 -0.069 -0.069
## F5 BY F2LOC2 28.542 -0.136 -0.107 -0.107
## F6 BY BYLOC2 11.385 0.131 0.084 0.084
## F6 BY F1CONF1 16.368 -0.139 -0.089 -0.089
## F6 BY F1LOC2 19.236 0.196 0.126 0.126
## F6 BY F2CONF1 15.535 -0.141 -0.090 -0.090
## F6 BY F2CONF3 14.912 -0.127 -0.081 -0.081
## F6 BY F2CONF4 69.947 0.281 0.180 0.180
##
##
## Beginning Time: 12:19:53
## Ending Time: 12:19:57
## Elapsed Time: 00:00:04
##
##
##
## MUTHEN & MUTHEN
## 3463 Stoner Ave.
## Los Angeles, CA 90066
##
## Tel: (310) 391-9971
## Fax: (310) 391-8971
## Web: www.StatModel.com
## Support: Support@StatModel.com
##
## Copyright (c) 1998-2019 Muthen & Muthen