Chapter 4 Confirmatory Factor Analysis
4.1 Syntax - R - One-Factor CFA
4.1.1 Use a sample covariance matrix an input
4.1.1.1 use the default reference indicator to identify the latent factor
AVRS <- '
64.000
37.120 64.000
35.200 32.640 64.000
28.800 32.640 33.280 64.000'
onef.cov <- getCov(AVRS, names = c("PATTERN", "COPYING", "MATRICES", "PAPERCUT"))
# Specify and fit the one-factor model
onef.model <- ' F1 =~ PATTERN + COPYING + MATRICES + PAPERCUT'
onef.fit <- cfa(onef.model, sample.cov = onef.cov, sample.nobs = 313)
summary(onef.fit, fit.measures = TRUE, standardized = TRUE)## lavaan 0.6-8 ended normally after 40 iterations
##
## Estimator ML
## Optimization method NLMINB
## Number of model parameters 8
##
## Number of observations 313
##
## Model Test User Model:
##
## Test statistic 7.405
## Degrees of freedom 2
## P-value (Chi-square) 0.025
##
## Model Test Baseline Model:
##
## Test statistic 405.864
## Degrees of freedom 6
## P-value 0.000
##
## User Model versus Baseline Model:
##
## Comparative Fit Index (CFI) 0.986
## Tucker-Lewis Index (TLI) 0.959
##
## Loglikelihood and Information Criteria:
##
## Loglikelihood user model (H0) -4178.740
## Loglikelihood unrestricted model (H1) -4175.037
##
## Akaike (AIC) 8373.479
## Bayesian (BIC) 8403.449
## Sample-size adjusted Bayesian (BIC) 8378.075
##
## Root Mean Square Error of Approximation:
##
## RMSEA 0.093
## 90 Percent confidence interval - lower 0.028
## 90 Percent confidence interval - upper 0.169
## P-value RMSEA <= 0.05 0.117
##
## Standardized Root Mean Square Residual:
##
## SRMR 0.023
##
## Parameter Estimates:
##
## Standard errors Standard
## Information Expected
## Information saturated (h1) model Structured
##
## Latent Variables:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## F1 =~
## PATTERN 1.000 5.940 0.744
## COPYING 1.006 0.089 11.364 0.000 5.978 0.748
## MATRICES 0.978 0.088 11.145 0.000 5.808 0.727
## PAPERCUT 0.895 0.086 10.368 0.000 5.318 0.666
##
## Variances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .PATTERN 28.512 3.230 8.828 0.000 28.512 0.447
## .COPYING 28.057 3.218 8.719 0.000 28.057 0.440
## .MATRICES 30.066 3.275 9.181 0.000 30.066 0.471
## .PAPERCUT 35.514 3.489 10.178 0.000 35.514 0.557
## F1 35.284 5.104 6.913 0.000 1.000 1.0004.1.1.2 use an alternate reference indicator to identify the latent factor
onef.model.altref <- ' F1 =~ NA*PATTERN + 1*COPYING + MATRICES + PAPERCUT'
onef.fit.altref <- cfa(onef.model.altref, sample.cov = onef.cov, sample.nobs = 313)
summary(onef.fit.altref, fit.measures = TRUE, standardized = TRUE)## lavaan 0.6-8 ended normally after 46 iterations
##
## Estimator ML
## Optimization method NLMINB
## Number of model parameters 8
##
## Number of observations 313
##
## Model Test User Model:
##
## Test statistic 7.405
## Degrees of freedom 2
## P-value (Chi-square) 0.025
##
## Model Test Baseline Model:
##
## Test statistic 405.864
## Degrees of freedom 6
## P-value 0.000
##
## User Model versus Baseline Model:
##
## Comparative Fit Index (CFI) 0.986
## Tucker-Lewis Index (TLI) 0.959
##
## Loglikelihood and Information Criteria:
##
## Loglikelihood user model (H0) -4178.740
## Loglikelihood unrestricted model (H1) -4175.037
##
## Akaike (AIC) 8373.479
## Bayesian (BIC) 8403.449
## Sample-size adjusted Bayesian (BIC) 8378.075
##
## Root Mean Square Error of Approximation:
##
## RMSEA 0.093
## 90 Percent confidence interval - lower 0.028
## 90 Percent confidence interval - upper 0.169
## P-value RMSEA <= 0.05 0.117
##
## Standardized Root Mean Square Residual:
##
## SRMR 0.023
##
## Parameter Estimates:
##
## Standard errors Standard
## Information Expected
## Information saturated (h1) model Structured
##
## Latent Variables:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## F1 =~
## PATTERN 0.994 0.087 11.364 0.000 5.940 0.744
## COPYING 1.000 5.978 0.748
## MATRICES 0.971 0.087 11.190 0.000 5.808 0.727
## PAPERCUT 0.890 0.085 10.406 0.000 5.318 0.666
##
## Variances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .PATTERN 28.512 3.230 8.828 0.000 28.512 0.447
## .COPYING 28.057 3.218 8.719 0.000 28.057 0.440
## .MATRICES 30.066 3.275 9.181 0.000 30.066 0.471
## .PAPERCUT 35.514 3.489 10.178 0.000 35.514 0.557
## F1 35.739 5.128 6.969 0.000 1.000 1.0004.1.1.3 fix latent variance at 1 to identify the latent factor
onef.model.fixvar <- '
F1 =~ NA*PATTERN + COPYING + MATRICES + PAPERCUT
F1 ~~ 1*F1'
onef.fit.fixvar <- cfa(onef.model.fixvar, sample.cov = onef.cov, sample.nobs = 313)
summary(onef.fit.fixvar, fit.measures = TRUE, standardized = TRUE)## lavaan 0.6-8 ended normally after 15 iterations
##
## Estimator ML
## Optimization method NLMINB
## Number of model parameters 8
##
## Number of observations 313
##
## Model Test User Model:
##
## Test statistic 7.405
## Degrees of freedom 2
## P-value (Chi-square) 0.025
##
## Model Test Baseline Model:
##
## Test statistic 405.864
## Degrees of freedom 6
## P-value 0.000
##
## User Model versus Baseline Model:
##
## Comparative Fit Index (CFI) 0.986
## Tucker-Lewis Index (TLI) 0.959
##
## Loglikelihood and Information Criteria:
##
## Loglikelihood user model (H0) -4178.740
## Loglikelihood unrestricted model (H1) -4175.037
##
## Akaike (AIC) 8373.479
## Bayesian (BIC) 8403.449
## Sample-size adjusted Bayesian (BIC) 8378.075
##
## Root Mean Square Error of Approximation:
##
## RMSEA 0.093
## 90 Percent confidence interval - lower 0.028
## 90 Percent confidence interval - upper 0.169
## P-value RMSEA <= 0.05 0.117
##
## Standardized Root Mean Square Residual:
##
## SRMR 0.023
##
## Parameter Estimates:
##
## Standard errors Standard
## Information Expected
## Information saturated (h1) model Structured
##
## Latent Variables:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## F1 =~
## PATTERN 5.940 0.430 13.827 0.000 5.940 0.744
## COPYING 5.978 0.429 13.938 0.000 5.978 0.748
## MATRICES 5.808 0.432 13.444 0.000 5.808 0.727
## PAPERCUT 5.318 0.442 12.042 0.000 5.318 0.666
##
## Variances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## F1 1.000 1.000 1.000
## .PATTERN 28.512 3.230 8.828 0.000 28.512 0.447
## .COPYING 28.057 3.218 8.719 0.000 28.057 0.440
## .MATRICES 30.066 3.275 9.181 0.000 30.066 0.471
## .PAPERCUT 35.514 3.489 10.178 0.000 35.514 0.5574.1.1.4 another way to fix latent variance at 1
by adding the “std.lv = TRUE” argument to the cfa() function call
onef.model <- ' F1 =~ PATTERN + COPYING + MATRICES + PAPERCUT'
onef.fit.fixvar.2 <- cfa(onef.model, sample.cov = onef.cov, sample.nobs = 313, std.lv = TRUE)
summary(onef.fit.fixvar.2, fit.measures = TRUE, standardized = TRUE)## lavaan 0.6-8 ended normally after 15 iterations
##
## Estimator ML
## Optimization method NLMINB
## Number of model parameters 8
##
## Number of observations 313
##
## Model Test User Model:
##
## Test statistic 7.405
## Degrees of freedom 2
## P-value (Chi-square) 0.025
##
## Model Test Baseline Model:
##
## Test statistic 405.864
## Degrees of freedom 6
## P-value 0.000
##
## User Model versus Baseline Model:
##
## Comparative Fit Index (CFI) 0.986
## Tucker-Lewis Index (TLI) 0.959
##
## Loglikelihood and Information Criteria:
##
## Loglikelihood user model (H0) -4178.740
## Loglikelihood unrestricted model (H1) -4175.037
##
## Akaike (AIC) 8373.479
## Bayesian (BIC) 8403.449
## Sample-size adjusted Bayesian (BIC) 8378.075
##
## Root Mean Square Error of Approximation:
##
## RMSEA 0.093
## 90 Percent confidence interval - lower 0.028
## 90 Percent confidence interval - upper 0.169
## P-value RMSEA <= 0.05 0.117
##
## Standardized Root Mean Square Residual:
##
## SRMR 0.023
##
## Parameter Estimates:
##
## Standard errors Standard
## Information Expected
## Information saturated (h1) model Structured
##
## Latent Variables:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## F1 =~
## PATTERN 5.940 0.430 13.827 0.000 5.940 0.744
## COPYING 5.978 0.429 13.938 0.000 5.978 0.748
## MATRICES 5.808 0.432 13.444 0.000 5.808 0.727
## PAPERCUT 5.318 0.442 12.042 0.000 5.318 0.666
##
## Variances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .PATTERN 28.512 3.230 8.828 0.000 28.512 0.447
## .COPYING 28.057 3.218 8.719 0.000 28.057 0.440
## .MATRICES 30.066 3.275 9.181 0.000 30.066 0.471
## .PAPERCUT 35.514 3.489 10.178 0.000 35.514 0.557
## F1 1.000 1.000 1.0004.1.2 Calculate coefficient omega \({\omega _u}\)
use reliability() function from the semTools package
## F1
## alpha 0.8125000
## omega 0.8129938
## omega2 0.8129938
## omega3 0.8128802
## avevar 0.5213312When raw data are available, the ci.reliability() function from the MBESS package can be used to get bootstrapped CI.
4.2 Syntax - R - Two-Factor CFA
4.2.1 An exmaple
AVRS2F <- '
64.000
37.120 64.000
35.200 32.640 64.000
28.800 32.640 33.280 64.000
33.280 31.360 36.480 32.640 64.000
34.560 30.080 47.360 39.040 42.240 64.000
21.120 4.480 16.640 17.280 27.520 28.160 64.000'
twof.cov <- getCov(AVRS2F, names = c("PATTERN", "COPYING", "MATRICES", "PAPERCUT", "QUANT", "NUMBSER", "EQUATION"))
twof.model <- '
ABSVIS =~ PATTERN + COPYING + MATRICES + PAPERCUT
QUANTITA =~ QUANT + NUMBSER + EQUATION
ABSVIS ~~ QUANTITA'
twof.fit <- cfa (twof.model, sample.cov = twof.cov, sample.nobs = 313, std.lv = TRUE)
summary(twof.fit, fit.measures = TRUE, standardized = TRUE, rsquare = TRUE)## lavaan 0.6-8 ended normally after 23 iterations
##
## Estimator ML
## Optimization method NLMINB
## Number of model parameters 15
##
## Number of observations 313
##
## Model Test User Model:
##
## Test statistic 110.322
## Degrees of freedom 13
## P-value (Chi-square) 0.000
##
## Model Test Baseline Model:
##
## Test statistic 1047.682
## Degrees of freedom 21
## P-value 0.000
##
## User Model versus Baseline Model:
##
## Comparative Fit Index (CFI) 0.905
## Tucker-Lewis Index (TLI) 0.847
##
## Loglikelihood and Information Criteria:
##
## Loglikelihood user model (H0) -7192.765
## Loglikelihood unrestricted model (H1) -7137.604
##
## Akaike (AIC) 14415.530
## Bayesian (BIC) 14471.723
## Sample-size adjusted Bayesian (BIC) 14424.148
##
## Root Mean Square Error of Approximation:
##
## RMSEA 0.155
## 90 Percent confidence interval - lower 0.129
## 90 Percent confidence interval - upper 0.182
## P-value RMSEA <= 0.05 0.000
##
## Standardized Root Mean Square Residual:
##
## SRMR 0.060
##
## Parameter Estimates:
##
## Standard errors Standard
## Information Expected
## Information saturated (h1) model Structured
##
## Latent Variables:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## ABSVIS =~
## PATTERN 5.532 0.416 13.308 0.000 5.532 0.693
## COPYING 5.153 0.425 12.135 0.000 5.153 0.645
## MATRICES 6.514 0.390 16.684 0.000 6.514 0.816
## PAPERCUT 5.530 0.416 13.301 0.000 5.530 0.692
## QUANTITA =~
## QUANT 6.055 0.401 15.108 0.000 6.055 0.758
## NUMBSER 7.160 0.376 19.047 0.000 7.160 0.896
## EQUATION 3.670 0.451 8.146 0.000 3.670 0.460
##
## Covariances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## ABSVIS ~~
## QUANTITA 0.938 0.024 39.281 0.000 0.938 0.938
##
## Variances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .PATTERN 33.191 3.037 10.927 0.000 33.191 0.520
## .COPYING 37.245 3.297 11.296 0.000 37.245 0.584
## .MATRICES 21.357 2.389 8.941 0.000 21.357 0.335
## .PAPERCUT 33.215 3.039 10.930 0.000 33.215 0.521
## .QUANT 27.134 2.638 10.287 0.000 27.134 0.425
## .NUMBSER 12.526 2.232 5.612 0.000 12.526 0.196
## .EQUATION 50.326 4.159 12.101 0.000 50.326 0.789
## ABSVIS 1.000 1.000 1.000
## QUANTITA 1.000 1.000 1.000
##
## R-Square:
## Estimate
## PATTERN 0.480
## COPYING 0.416
## MATRICES 0.665
## PAPERCUT 0.479
## QUANT 0.575
## NUMBSER 0.804
## EQUATION 0.211Obtain model-implied (fitted) covariance matrix and mean vector
## $cov
## PATTER COPYIN MATRIC PAPERC QUANT NUMBSE EQUATI
## PATTERN 63.796
## COPYING 28.505 63.796
## MATRICES 36.039 33.567 63.796
## PAPERCUT 30.593 28.494 36.025 63.796
## QUANT 31.435 29.279 37.017 31.423 63.796
## NUMBSER 37.174 34.624 43.775 37.159 43.355 63.796
## EQUATION 19.054 17.747 22.438 19.047 22.222 26.279 63.796Obtain unstandardized residuals of a fitted model
## $type
## [1] "raw"
##
## $cov
## PATTER COPYIN MATRIC PAPERC QUANT NUMBSE EQUATI
## PATTERN 0.000
## COPYING 8.496 0.000
## MATRICES -0.952 -1.031 0.000
## PAPERCUT -1.885 4.042 -2.851 0.000
## QUANT 1.738 1.981 -0.654 1.113 0.000
## NUMBSER -2.724 -4.640 3.434 1.756 -1.250 0.000
## EQUATION 1.998 -13.281 -5.851 -1.822 5.210 1.791 0.000Obtain standardized residuals of a fitted model
## $type
## [1] "standardized"
##
## $cov
## PATTER COPYIN MATRIC PAPERC QUANT
## PATTERN -1.215870e+09
## COPYING 4.439000e+00 0.000000e+00
## MATRICES -9.220000e-01 -9.230000e-01 -1.606608e+09
## PAPERCUT -1.239000e+00 2.247000e+00 -2.824000e+00 -1.249981e+09
## QUANT 1.195000e+00 1.242000e+00 -6.040000e-01 7.320000e-01 -1.116695e+09
## NUMBSER -3.145000e+00 -5.016000e+00 4.763000e+00 1.737000e+00 -4.772000e+00
## EQUATION 8.640000e-01 -5.349000e+00 -3.404000e+00 -8.080000e-01 2.660000e+00
## NUMBSE EQUATI
## PATTERN
## COPYING
## MATRICES
## PAPERCUT
## QUANT
## NUMBSER -2.506552e+09
## EQUATION 1.918000e+00 -6.401233e+09Request a list of model matrices counting free parameters in the model
## $lambda
## ABSVIS QUANTI
## PATTERN 1 0
## COPYING 2 0
## MATRICES 3 0
## PAPERCUT 4 0
## QUANT 0 5
## NUMBSER 0 6
## EQUATION 0 7
##
## $theta
## PATTER COPYIN MATRIC PAPERC QUANT NUMBSE EQUATI
## PATTERN 9
## COPYING 0 10
## MATRICES 0 0 11
## PAPERCUT 0 0 0 12
## QUANT 0 0 0 0 13
## NUMBSER 0 0 0 0 0 14
## EQUATION 0 0 0 0 0 0 15
##
## $psi
## ABSVIS QUANTI
## ABSVIS 0
## QUANTITA 8 0Plot the path diagram
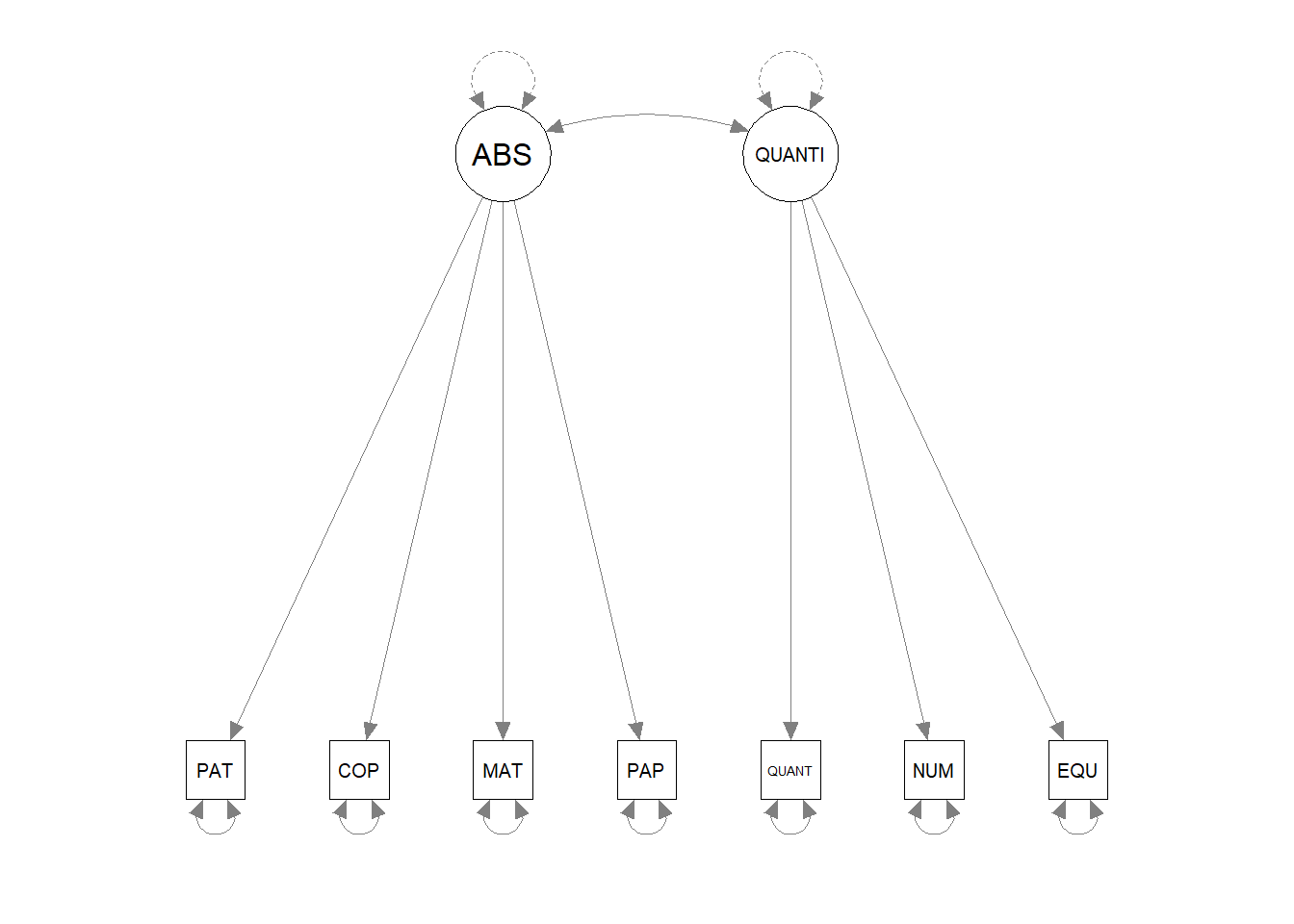
4.2.2 Model comparison example - two-factor CFA model across time
WERHLE <- '
.903
1.522 7.126
.508 1.577 .933
.387 1.081 .527 1.117
1.036 3.499 1.363 1.715 6.065
.441 1.404 .674 .649 1.742 .974'
twof.acrosstime.cov <- getCov(WERHLE, names = c("CHANGE1", "INFLUEN1", "SUGGEST1", "CHANGE2", "INFLUEN2", "SUGGEST2"))
twof.acrosstime.model1 <- '
CONTROL1 =~ CHANGE1 + INFLUEN1 + SUGGEST1
CONTROL2 =~ CHANGE2 + INFLUEN2 + SUGGEST2'
twof.acrosstime.model2 <- '
CONTROL1 =~ CHANGE1 + INFLUEN1 + SUGGEST1
CONTROL2 =~ CHANGE2 + INFLUEN2 + SUGGEST2
CHANGE1 ~~ CHANGE2
INFLUEN1 ~~ INFLUEN2
SUGGEST1 ~~ SUGGEST2'
twof.acrosstime.model3 <- '
CONTROL1 =~ a*CHANGE1 + b*INFLUEN1 + c*SUGGEST1
CONTROL2 =~ a*CHANGE2 + b*INFLUEN2 + c*SUGGEST2'
twof.acrosstime.model4 <- '
CONTROL1 =~ a*CHANGE1 + b*INFLUEN1 + c*SUGGEST1
CONTROL2 =~ a*CHANGE2 + b*INFLUEN2 + c*SUGGEST2
CHANGE1 ~~ CHANGE2
INFLUEN1 ~~ INFLUEN2
SUGGEST1 ~~ SUGGEST2'
twof.acrosstime.fit1 <- cfa(twof.acrosstime.model1, sample.cov = twof.acrosstime.cov, sample.nobs = 119, std.lv = TRUE)
twof.acrosstime.fit2 <- cfa(twof.acrosstime.model2, sample.cov = twof.acrosstime.cov, sample.nobs = 119, std.lv = TRUE)
twof.acrosstime.fit3 <- cfa(twof.acrosstime.model3, sample.cov = twof.acrosstime.cov, sample.nobs = 119, std.lv = TRUE)
twof.acrosstime.fit4 <- cfa(twof.acrosstime.model4, sample.cov = twof.acrosstime.cov, sample.nobs = 119, std.lv = TRUE)
summary(twof.acrosstime.fit1, fit.measures = TRUE, standardized = TRUE, rsquare = TRUE)## lavaan 0.6-8 ended normally after 20 iterations
##
## Estimator ML
## Optimization method NLMINB
## Number of model parameters 13
##
## Number of observations 119
##
## Model Test User Model:
##
## Test statistic 18.648
## Degrees of freedom 8
## P-value (Chi-square) 0.017
##
## Model Test Baseline Model:
##
## Test statistic 381.003
## Degrees of freedom 15
## P-value 0.000
##
## User Model versus Baseline Model:
##
## Comparative Fit Index (CFI) 0.971
## Tucker-Lewis Index (TLI) 0.945
##
## Loglikelihood and Information Criteria:
##
## Loglikelihood user model (H0) -1047.844
## Loglikelihood unrestricted model (H1) -1038.520
##
## Akaike (AIC) 2121.689
## Bayesian (BIC) 2157.817
## Sample-size adjusted Bayesian (BIC) 2116.719
##
## Root Mean Square Error of Approximation:
##
## RMSEA 0.106
## 90 Percent confidence interval - lower 0.042
## 90 Percent confidence interval - upper 0.169
## P-value RMSEA <= 0.05 0.068
##
## Standardized Root Mean Square Residual:
##
## SRMR 0.038
##
## Parameter Estimates:
##
## Standard errors Standard
## Information Expected
## Information saturated (h1) model Structured
##
## Latent Variables:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## CONTROL1 =~
## CHANGE1 0.645 0.081 7.924 0.000 0.645 0.681
## INFLUEN1 1.985 0.222 8.950 0.000 1.985 0.747
## SUGGEST1 0.815 0.077 10.630 0.000 0.815 0.847
## CONTROL2 =~
## CHANGE2 0.769 0.087 8.857 0.000 0.769 0.731
## INFLUEN2 2.039 0.192 10.638 0.000 2.039 0.831
## SUGGEST2 0.861 0.075 11.486 0.000 0.861 0.876
##
## Covariances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## CONTROL1 ~~
## CONTROL2 0.846 0.047 18.040 0.000 0.846 0.846
##
## Variances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .CHANGE1 0.480 0.073 6.614 0.000 0.480 0.536
## .INFLUEN1 3.126 0.514 6.079 0.000 3.126 0.442
## .SUGGEST1 0.261 0.059 4.409 0.000 0.261 0.282
## .CHANGE2 0.517 0.079 6.578 0.000 0.517 0.466
## .INFLUEN2 1.856 0.345 5.383 0.000 1.856 0.309
## .SUGGEST2 0.225 0.052 4.371 0.000 0.225 0.233
## CONTROL1 1.000 1.000 1.000
## CONTROL2 1.000 1.000 1.000
##
## R-Square:
## Estimate
## CHANGE1 0.464
## INFLUEN1 0.558
## SUGGEST1 0.718
## CHANGE2 0.534
## INFLUEN2 0.691
## SUGGEST2 0.767## lavaan 0.6-8 ended normally after 26 iterations
##
## Estimator ML
## Optimization method NLMINB
## Number of model parameters 16
##
## Number of observations 119
##
## Model Test User Model:
##
## Test statistic 7.750
## Degrees of freedom 5
## P-value (Chi-square) 0.171
##
## Model Test Baseline Model:
##
## Test statistic 381.003
## Degrees of freedom 15
## P-value 0.000
##
## User Model versus Baseline Model:
##
## Comparative Fit Index (CFI) 0.992
## Tucker-Lewis Index (TLI) 0.977
##
## Loglikelihood and Information Criteria:
##
## Loglikelihood user model (H0) -1042.395
## Loglikelihood unrestricted model (H1) -1038.520
##
## Akaike (AIC) 2116.791
## Bayesian (BIC) 2161.257
## Sample-size adjusted Bayesian (BIC) 2110.675
##
## Root Mean Square Error of Approximation:
##
## RMSEA 0.068
## 90 Percent confidence interval - lower 0.000
## 90 Percent confidence interval - upper 0.156
## P-value RMSEA <= 0.05 0.310
##
## Standardized Root Mean Square Residual:
##
## SRMR 0.029
##
## Parameter Estimates:
##
## Standard errors Standard
## Information Expected
## Information saturated (h1) model Structured
##
## Latent Variables:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## CONTROL1 =~
## CHANGE1 0.672 0.082 8.185 0.000 0.672 0.710
## INFLUEN1 2.051 0.226 9.058 0.000 2.051 0.771
## SUGGEST1 0.765 0.079 9.703 0.000 0.765 0.800
## CONTROL2 =~
## CHANGE2 0.782 0.087 8.952 0.000 0.782 0.744
## INFLUEN2 2.107 0.193 10.903 0.000 2.107 0.859
## SUGGEST2 0.813 0.077 10.585 0.000 0.813 0.835
##
## Covariances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .CHANGE1 ~~
## .CHANGE2 -0.016 0.053 -0.301 0.763 -0.016 -0.034
## .INFLUEN1 ~~
## .INFLUEN2 0.092 0.310 0.296 0.767 0.092 0.043
## .SUGGEST1 ~~
## .SUGGEST2 0.130 0.046 2.860 0.004 0.130 0.424
## CONTROL1 ~~
## CONTROL2 0.810 0.050 16.247 0.000 0.810 0.810
##
## Variances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .CHANGE1 0.444 0.073 6.089 0.000 0.444 0.496
## .INFLUEN1 2.864 0.547 5.236 0.000 2.864 0.405
## .SUGGEST1 0.328 0.066 4.947 0.000 0.328 0.359
## .CHANGE2 0.495 0.080 6.222 0.000 0.495 0.447
## .INFLUEN2 1.579 0.373 4.239 0.000 1.579 0.262
## .SUGGEST2 0.288 0.059 4.925 0.000 0.288 0.304
## CONTROL1 1.000 1.000 1.000
## CONTROL2 1.000 1.000 1.000
##
## R-Square:
## Estimate
## CHANGE1 0.504
## INFLUEN1 0.595
## SUGGEST1 0.641
## CHANGE2 0.553
## INFLUEN2 0.738
## SUGGEST2 0.696## lavaan 0.6-8 ended normally after 17 iterations
##
## Estimator ML
## Optimization method NLMINB
## Number of model parameters 13
## Number of equality constraints 3
##
## Number of observations 119
##
## Model Test User Model:
##
## Test statistic 20.206
## Degrees of freedom 11
## P-value (Chi-square) 0.043
##
## Model Test Baseline Model:
##
## Test statistic 381.003
## Degrees of freedom 15
## P-value 0.000
##
## User Model versus Baseline Model:
##
## Comparative Fit Index (CFI) 0.975
## Tucker-Lewis Index (TLI) 0.966
##
## Loglikelihood and Information Criteria:
##
## Loglikelihood user model (H0) -1048.623
## Loglikelihood unrestricted model (H1) -1038.520
##
## Akaike (AIC) 2117.247
## Bayesian (BIC) 2145.038
## Sample-size adjusted Bayesian (BIC) 2113.424
##
## Root Mean Square Error of Approximation:
##
## RMSEA 0.084
## 90 Percent confidence interval - lower 0.015
## 90 Percent confidence interval - upper 0.141
## P-value RMSEA <= 0.05 0.154
##
## Standardized Root Mean Square Residual:
##
## SRMR 0.055
##
## Parameter Estimates:
##
## Standard errors Standard
## Information Expected
## Information saturated (h1) model Structured
##
## Latent Variables:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## CONTROL1 =~
## CHANGE1 (a) 0.706 0.065 10.787 0.000 0.706 0.717
## INFLUEN1 (b) 2.014 0.164 12.299 0.000 2.014 0.753
## SUGGEST1 (c) 0.838 0.063 13.395 0.000 0.838 0.854
## CONTROL2 =~
## CHANGE2 (a) 0.706 0.065 10.787 0.000 0.706 0.696
## INFLUEN2 (b) 2.014 0.164 12.299 0.000 2.014 0.829
## SUGGEST2 (c) 0.838 0.063 13.395 0.000 0.838 0.869
##
## Covariances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## CONTROL1 ~~
## CONTROL2 0.843 0.047 17.998 0.000 0.843 0.843
##
## Variances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .CHANGE1 0.471 0.072 6.516 0.000 0.471 0.486
## .INFLUEN1 3.107 0.498 6.242 0.000 3.107 0.434
## .SUGGEST1 0.261 0.056 4.630 0.000 0.261 0.271
## .CHANGE2 0.529 0.078 6.797 0.000 0.529 0.515
## .INFLUEN2 1.851 0.341 5.435 0.000 1.851 0.313
## .SUGGEST2 0.227 0.050 4.564 0.000 0.227 0.244
## CONTROL1 1.000 1.000 1.000
## CONTROL2 1.000 1.000 1.000
##
## R-Square:
## Estimate
## CHANGE1 0.514
## INFLUEN1 0.566
## SUGGEST1 0.729
## CHANGE2 0.485
## INFLUEN2 0.687
## SUGGEST2 0.756## lavaan 0.6-8 ended normally after 24 iterations
##
## Estimator ML
## Optimization method NLMINB
## Number of model parameters 16
## Number of equality constraints 3
##
## Number of observations 119
##
## Model Test User Model:
##
## Test statistic 9.022
## Degrees of freedom 8
## P-value (Chi-square) 0.340
##
## Model Test Baseline Model:
##
## Test statistic 381.003
## Degrees of freedom 15
## P-value 0.000
##
## User Model versus Baseline Model:
##
## Comparative Fit Index (CFI) 0.997
## Tucker-Lewis Index (TLI) 0.995
##
## Loglikelihood and Information Criteria:
##
## Loglikelihood user model (H0) -1043.031
## Loglikelihood unrestricted model (H1) -1038.520
##
## Akaike (AIC) 2112.062
## Bayesian (BIC) 2148.191
## Sample-size adjusted Bayesian (BIC) 2107.093
##
## Root Mean Square Error of Approximation:
##
## RMSEA 0.033
## 90 Percent confidence interval - lower 0.000
## 90 Percent confidence interval - upper 0.116
## P-value RMSEA <= 0.05 0.548
##
## Standardized Root Mean Square Residual:
##
## SRMR 0.048
##
## Parameter Estimates:
##
## Standard errors Standard
## Information Expected
## Information saturated (h1) model Structured
##
## Latent Variables:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## CONTROL1 =~
## CHANGE1 (a) 0.725 0.065 11.116 0.000 0.725 0.740
## INFLUEN1 (b) 2.081 0.168 12.415 0.000 2.081 0.776
## SUGGEST1 (c) 0.789 0.067 11.687 0.000 0.789 0.809
## CONTROL2 =~
## CHANGE2 (a) 0.725 0.065 11.116 0.000 0.725 0.714
## INFLUEN2 (b) 2.081 0.168 12.415 0.000 2.081 0.858
## SUGGEST2 (c) 0.789 0.067 11.687 0.000 0.789 0.825
##
## Covariances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .CHANGE1 ~~
## .CHANGE2 -0.018 0.053 -0.336 0.737 -0.018 -0.038
## .INFLUEN1 ~~
## .INFLUEN2 0.085 0.309 0.274 0.784 0.085 0.040
## .SUGGEST1 ~~
## .SUGGEST2 0.132 0.046 2.897 0.004 0.132 0.428
## CONTROL1 ~~
## CONTROL2 0.807 0.050 16.183 0.000 0.807 0.807
##
## Variances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .CHANGE1 0.434 0.072 6.016 0.000 0.434 0.452
## .INFLUEN1 2.862 0.521 5.492 0.000 2.862 0.398
## .SUGGEST1 0.328 0.064 5.080 0.000 0.328 0.345
## .CHANGE2 0.507 0.078 6.477 0.000 0.507 0.491
## .INFLUEN2 1.558 0.367 4.243 0.000 1.558 0.265
## .SUGGEST2 0.292 0.058 5.061 0.000 0.292 0.319
## CONTROL1 1.000 1.000 1.000
## CONTROL2 1.000 1.000 1.000
##
## R-Square:
## Estimate
## CHANGE1 0.548
## INFLUEN1 0.602
## SUGGEST1 0.655
## CHANGE2 0.509
## INFLUEN2 0.735
## SUGGEST2 0.681Use the anova() function or the lavTestLRT() function from lavaan to compare nested models
## Chi-Squared Difference Test
##
## Df AIC BIC Chisq Chisq diff Df diff Pr(>Chisq)
## twof.acrosstime.fit2 5 2116.8 2161.3 7.750
## twof.acrosstime.fit1 8 2121.7 2157.8 18.648 10.898 3 0.01229 *
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1## Chi-Squared Difference Test
##
## Df AIC BIC Chisq Chisq diff Df diff Pr(>Chisq)
## twof.acrosstime.fit2 5 2116.8 2161.3 7.750
## twof.acrosstime.fit1 8 2121.7 2157.8 18.648 10.898 3 0.01229 *
## ---
## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1Plot the path diagram
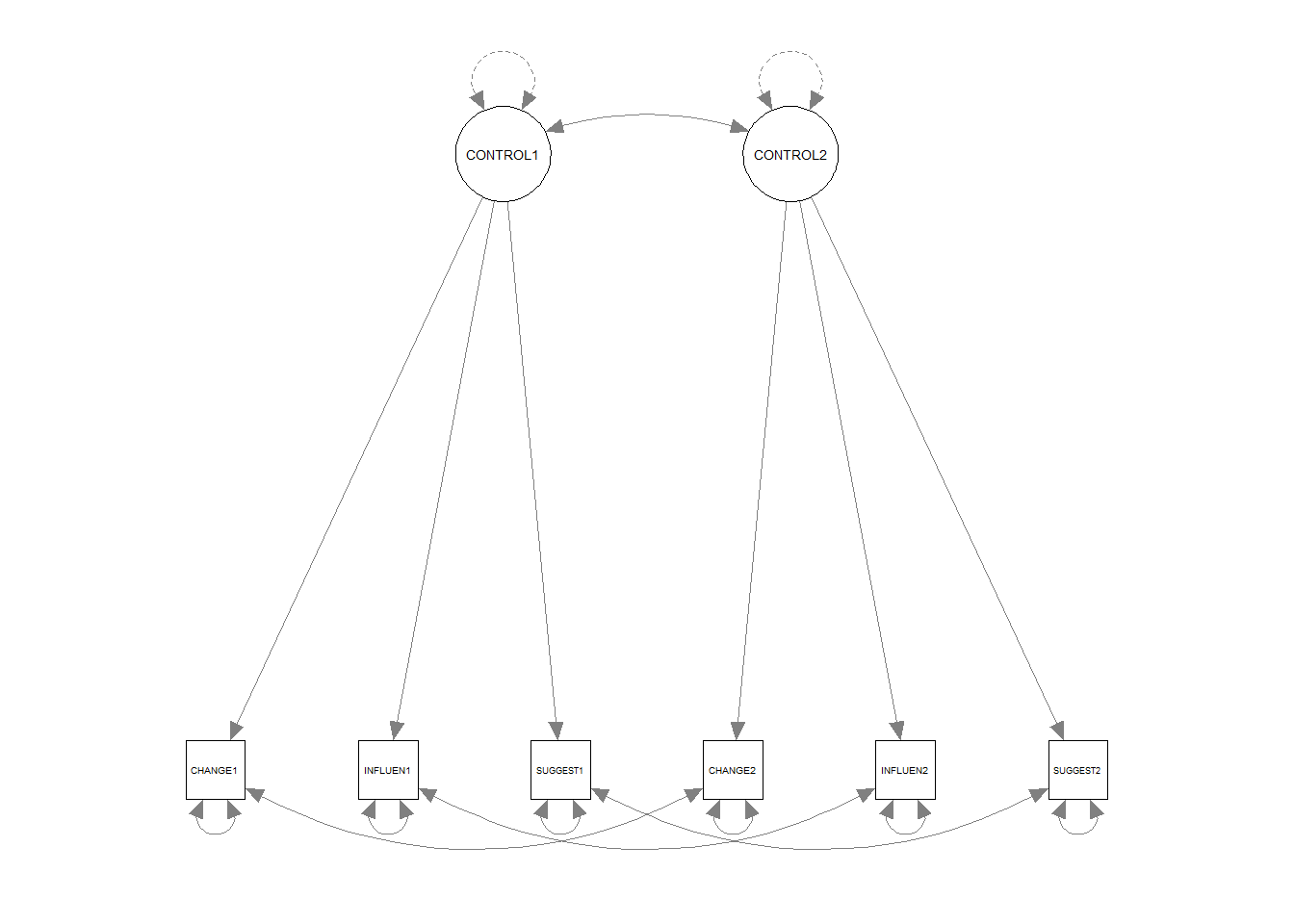
4.3 Syntax - Mplus - One-Factor CFA
4.3.1 Use a sample covariance matrix an input
4.3.1.1 use the default reference indicator to identify the latent factor
TITLE: ONE FACTOR CONFIRMATORY FACTOR ANALYSIS;
ABSTRACT VISUAL REASONING SCALE - SB4
DATA: FILE IS "data\AVRS.DAT";
TYPE IS COVARIANCE;
NOBSERVATIONS ARE 313;
VARIABLE: NAMES ARE PATTERN COPYING MATRICES PAPERCUT;
MODEL: F1 BY PATTERN COPYING MATRICES PAPERCUT;
OUTPUT: SAMPSTAT STANDARDIZED(STDYX)RESIDUAL;## Mplus VERSION 8.4
## MUTHEN & MUTHEN
## 06/10/2021 12:19 PM
##
## INPUT INSTRUCTIONS
##
## TITLE: ONE FACTOR CONFIRMATORY FACTOR ANALYSIS;
## ABSTRACT VISUAL REASONING SCALE - SB4
## DATA: FILE IS "data\AVRS.DAT";
## TYPE IS COVARIANCE;
## NOBSERVATIONS ARE 313;
## VARIABLE: NAMES ARE PATTERN COPYING MATRICES PAPERCUT;
## MODEL: F1 BY PATTERN COPYING MATRICES PAPERCUT;
## OUTPUT: SAMPSTAT STANDARDIZED(STDYX)RESIDUAL;
##
##
##
## 1 ERROR(S) FOUND IN THE INPUT INSTRUCTIONS
##
##
##
## ONE FACTOR CONFIRMATORY FACTOR ANALYSIS;
## ABSTRACT VISUAL REASONING SCALE - SB4
##
## SUMMARY OF ANALYSIS
##
## Number of groups 1
## Number of observations 313
##
## Number of dependent variables 4
## Number of independent variables 0
## Number of continuous latent variables 1
##
## Observed dependent variables
##
## Continuous
## PATTERN COPYING MATRICES PAPERCUT
##
## Continuous latent variables
## F1
##
##
## Estimator ML
## Information matrix EXPECTED
## Maximum number of iterations 1000
## Convergence criterion 0.500D-04
## Maximum number of steepest descent iterations 20
##
## Input data file(s)
## data\AVRS.DAT
##
## Input data format FREE
##
##
## SAMPLE STATISTICS
##
##
## SAMPLE STATISTICS
##
##
## Covariances/Correlations/Residual Correlations
## PATTERN COPYING MATRICES PAPERCUT
## ________ ________ ________ ________
## PATTERN 64.000
## COPYING 37.120 64.000
## MATRICES 35.200 32.640 64.000
## PAPERCUT 28.800 32.640 33.280 64.000
##
##
## THE MODEL ESTIMATION TERMINATED NORMALLY
##
##
##
## MODEL FIT INFORMATION
##
## Number of Free Parameters 8
##
## Loglikelihood
##
## H0 Value -4178.740
## H1 Value -4175.037
##
## Information Criteria
##
## Akaike (AIC) 8373.479
## Bayesian (BIC) 8403.449
## Sample-Size Adjusted BIC 8378.075
## (n* = (n + 2) / 24)
##
## Chi-Square Test of Model Fit
##
## Value 7.405
## Degrees of Freedom 2
## P-Value 0.0247
##
## RMSEA (Root Mean Square Error Of Approximation)
##
## Estimate 0.093
## 90 Percent C.I. 0.028 0.169
## Probability RMSEA <= .05 0.117
##
## CFI/TLI
##
## CFI 0.986
## TLI 0.959
##
## Chi-Square Test of Model Fit for the Baseline Model
##
## Value 405.864
## Degrees of Freedom 6
## P-Value 0.0000
##
## SRMR (Standardized Root Mean Square Residual)
##
## Value 0.023
##
##
##
## MODEL RESULTS
##
## Two-Tailed
## Estimate S.E. Est./S.E. P-Value
##
## F1 BY
## PATTERN 1.000 0.000 999.000 999.000
## COPYING 1.006 0.089 11.364 0.000
## MATRICES 0.978 0.088 11.145 0.000
## PAPERCUT 0.895 0.086 10.368 0.000
##
## Variances
## F1 35.283 5.104 6.913 0.000
##
## Residual Variances
## PATTERN 28.511 3.230 8.828 0.000
## COPYING 28.057 3.218 8.719 0.000
## MATRICES 30.065 3.275 9.181 0.000
## PAPERCUT 35.514 3.489 10.178 0.000
##
##
## STANDARDIZED MODEL RESULTS
##
##
## STDYX Standardization
##
## Two-Tailed
## Estimate S.E. Est./S.E. P-Value
##
## F1 BY
## PATTERN 0.744 0.035 21.295 0.000
## COPYING 0.748 0.035 21.590 0.000
## MATRICES 0.727 0.036 20.288 0.000
## PAPERCUT 0.666 0.040 16.855 0.000
##
## Variances
## F1 1.000 0.000 999.000 999.000
##
## Residual Variances
## PATTERN 0.447 0.052 8.604 0.000
## COPYING 0.440 0.052 8.475 0.000
## MATRICES 0.471 0.052 9.042 0.000
## PAPERCUT 0.557 0.053 10.583 0.000
##
##
## R-SQUARE
##
## Observed Two-Tailed
## Variable Estimate S.E. Est./S.E. P-Value
##
## PATTERN 0.553 0.052 10.647 0.000
## COPYING 0.560 0.052 10.795 0.000
## MATRICES 0.529 0.052 10.144 0.000
## PAPERCUT 0.443 0.053 8.428 0.000
##
##
## QUALITY OF NUMERICAL RESULTS
##
## Condition Number for the Information Matrix 0.302E-01
## (ratio of smallest to largest eigenvalue)
##
##
## RESIDUAL OUTPUT
##
##
## ESTIMATED MODEL AND RESIDUALS (OBSERVED - ESTIMATED)
##
##
## Model Estimated Covariances/Correlations/Residual Correlations
## PATTERN COPYING MATRICES PAPERCUT
## ________ ________ ________ ________
## PATTERN 63.794
## COPYING 35.510 63.796
## MATRICES 34.498 34.720 63.796
## PAPERCUT 31.589 31.792 30.886 63.796
##
##
## Residuals for Covariances/Correlations/Residual Correlations
## PATTERN COPYING MATRICES PAPERCUT
## ________ ________ ________ ________
## PATTERN 0.002
## COPYING 1.491 0.000
## MATRICES 0.589 -2.184 0.000
## PAPERCUT -2.881 0.744 2.288 0.000
##
##
## Standardized Residuals (z-scores) for Covariances/Correlations/Residual Corr
## PATTERN COPYING MATRICES PAPERCUT
## ________ ________ ________ ________
## PATTERN 0.043
## COPYING 1.516 999.000
## MATRICES 0.622 -4.270 999.000
## PAPERCUT -3.400 0.635 1.646 999.000
##
##
## Normalized Residuals for Covariances/Correlations/Residual Correlations
## PATTERN COPYING MATRICES PAPERCUT
## ________ ________ ________ ________
## PATTERN 0.000
## COPYING 0.358 0.000
## MATRICES 0.143 -0.540 0.000
## PAPERCUT -0.729 0.184 0.563 0.000
##
##
## Beginning Time: 12:19:13
## Ending Time: 12:19:13
## Elapsed Time: 00:00:00
##
##
##
## MUTHEN & MUTHEN
## 3463 Stoner Ave.
## Los Angeles, CA 90066
##
## Tel: (310) 391-9971
## Fax: (310) 391-8971
## Web: www.StatModel.com
## Support: Support@StatModel.com
##
## Copyright (c) 1998-2019 Muthen & Muthen4.3.1.2 use an alternate reference indicator to identify the latent factor
TITLE: ONE FACTOR CONFIRMATORY FACTOR ANALYSIS;
ABSTRACT VISUAL REASONING SCALE - SB4
DATA: FILE IS "data\AVRS.DAT";
TYPE IS COVARIANCE;
NOBSERVATIONS ARE 313;
VARIABLE: NAMES ARE PATTERN COPYING MATRICES PAPERCUT;
MODEL: F1 BY PATTERN* COPYING@1 MATRICES PAPERCUT;
OUTPUT: SAMPSTAT STANDARDIZED(STDYX) RESIDUAL;## Mplus VERSION 8.4
## MUTHEN & MUTHEN
## 06/10/2021 12:19 PM
##
## INPUT INSTRUCTIONS
##
## TITLE: ONE FACTOR CONFIRMATORY FACTOR ANALYSIS;
## ABSTRACT VISUAL REASONING SCALE - SB4
## DATA: FILE IS "data\AVRS.DAT";
## TYPE IS COVARIANCE;
## NOBSERVATIONS ARE 313;
## VARIABLE: NAMES ARE PATTERN COPYING MATRICES PAPERCUT;
## MODEL: F1 BY PATTERN* COPYING@1 MATRICES PAPERCUT;
## OUTPUT: SAMPSTAT STANDARDIZED(STDYX) RESIDUAL;
##
##
##
## 1 ERROR(S) FOUND IN THE INPUT INSTRUCTIONS
##
##
##
## ONE FACTOR CONFIRMATORY FACTOR ANALYSIS;
## ABSTRACT VISUAL REASONING SCALE - SB4
##
## SUMMARY OF ANALYSIS
##
## Number of groups 1
## Number of observations 313
##
## Number of dependent variables 4
## Number of independent variables 0
## Number of continuous latent variables 1
##
## Observed dependent variables
##
## Continuous
## PATTERN COPYING MATRICES PAPERCUT
##
## Continuous latent variables
## F1
##
##
## Estimator ML
## Information matrix EXPECTED
## Maximum number of iterations 1000
## Convergence criterion 0.500D-04
## Maximum number of steepest descent iterations 20
##
## Input data file(s)
## data\AVRS.DAT
##
## Input data format FREE
##
##
## SAMPLE STATISTICS
##
##
## SAMPLE STATISTICS
##
##
## Covariances/Correlations/Residual Correlations
## PATTERN COPYING MATRICES PAPERCUT
## ________ ________ ________ ________
## PATTERN 64.000
## COPYING 37.120 64.000
## MATRICES 35.200 32.640 64.000
## PAPERCUT 28.800 32.640 33.280 64.000
##
##
## THE MODEL ESTIMATION TERMINATED NORMALLY
##
##
##
## MODEL FIT INFORMATION
##
## Number of Free Parameters 8
##
## Loglikelihood
##
## H0 Value -4178.740
## H1 Value -4175.037
##
## Information Criteria
##
## Akaike (AIC) 8373.479
## Bayesian (BIC) 8403.449
## Sample-Size Adjusted BIC 8378.075
## (n* = (n + 2) / 24)
##
## Chi-Square Test of Model Fit
##
## Value 7.405
## Degrees of Freedom 2
## P-Value 0.0247
##
## RMSEA (Root Mean Square Error Of Approximation)
##
## Estimate 0.093
## 90 Percent C.I. 0.028 0.169
## Probability RMSEA <= .05 0.117
##
## CFI/TLI
##
## CFI 0.986
## TLI 0.959
##
## Chi-Square Test of Model Fit for the Baseline Model
##
## Value 405.864
## Degrees of Freedom 6
## P-Value 0.0000
##
## SRMR (Standardized Root Mean Square Residual)
##
## Value 0.023
##
##
##
## MODEL RESULTS
##
## Two-Tailed
## Estimate S.E. Est./S.E. P-Value
##
## F1 BY
## PATTERN 0.994 0.087 11.364 0.000
## COPYING 1.000 0.000 999.000 999.000
## MATRICES 0.972 0.087 11.190 0.000
## PAPERCUT 0.890 0.085 10.406 0.000
##
## Variances
## F1 35.738 5.128 6.969 0.000
##
## Residual Variances
## PATTERN 28.511 3.230 8.828 0.000
## COPYING 28.058 3.218 8.719 0.000
## MATRICES 30.065 3.275 9.181 0.000
## PAPERCUT 35.514 3.489 10.178 0.000
##
##
## STANDARDIZED MODEL RESULTS
##
##
## STDYX Standardization
##
## Two-Tailed
## Estimate S.E. Est./S.E. P-Value
##
## F1 BY
## PATTERN 0.744 0.035 21.295 0.000
## COPYING 0.748 0.035 21.590 0.000
## MATRICES 0.727 0.036 20.288 0.000
## PAPERCUT 0.666 0.040 16.855 0.000
##
## Variances
## F1 1.000 0.000 999.000 999.000
##
## Residual Variances
## PATTERN 0.447 0.052 8.604 0.000
## COPYING 0.440 0.052 8.475 0.000
## MATRICES 0.471 0.052 9.042 0.000
## PAPERCUT 0.557 0.053 10.583 0.000
##
##
## R-SQUARE
##
## Observed Two-Tailed
## Variable Estimate S.E. Est./S.E. P-Value
##
## PATTERN 0.553 0.052 10.647 0.000
## COPYING 0.560 0.052 10.795 0.000
## MATRICES 0.529 0.052 10.144 0.000
## PAPERCUT 0.443 0.053 8.428 0.000
##
##
## QUALITY OF NUMERICAL RESULTS
##
## Condition Number for the Information Matrix 0.303E-01
## (ratio of smallest to largest eigenvalue)
##
##
## RESIDUAL OUTPUT
##
##
## ESTIMATED MODEL AND RESIDUALS (OBSERVED - ESTIMATED)
##
##
## Model Estimated Covariances/Correlations/Residual Correlations
## PATTERN COPYING MATRICES PAPERCUT
## ________ ________ ________ ________
## PATTERN 63.796
## COPYING 35.510 63.796
## MATRICES 34.498 34.720 63.795
## PAPERCUT 31.589 31.792 30.885 63.795
##
##
## Residuals for Covariances/Correlations/Residual Correlations
## PATTERN COPYING MATRICES PAPERCUT
## ________ ________ ________ ________
## PATTERN 0.000
## COPYING 1.491 0.000
## MATRICES 0.589 -2.184 0.001
## PAPERCUT -2.881 0.744 2.288 0.001
##
##
## Standardized Residuals (z-scores) for Covariances/Correlations/Residual Corr
## PATTERN COPYING MATRICES PAPERCUT
## ________ ________ ________ ________
## PATTERN 0.005
## COPYING 1.516 999.000
## MATRICES 0.622 -4.267 0.026
## PAPERCUT -3.400 0.635 1.646 0.035
##
##
## Normalized Residuals for Covariances/Correlations/Residual Correlations
## PATTERN COPYING MATRICES PAPERCUT
## ________ ________ ________ ________
## PATTERN 0.000
## COPYING 0.358 0.000
## MATRICES 0.143 -0.540 0.000
## PAPERCUT -0.729 0.184 0.563 0.000
##
##
## Beginning Time: 12:19:13
## Ending Time: 12:19:13
## Elapsed Time: 00:00:00
##
##
##
## MUTHEN & MUTHEN
## 3463 Stoner Ave.
## Los Angeles, CA 90066
##
## Tel: (310) 391-9971
## Fax: (310) 391-8971
## Web: www.StatModel.com
## Support: Support@StatModel.com
##
## Copyright (c) 1998-2019 Muthen & Muthen4.3.1.3 fix latent variance at 1 to identify the latent factor
TITLE: ONE FACTOR CONFIRMATORY FACTOR ANALYSIS;
ABSTRACT VISUAL REASONING SCALE - SB4
DATA: FILE IS "data\AVRS.DAT";
TYPE IS COVARIANCE;
NOBSERVATIONS ARE 313;
VARIABLE: NAMES ARE PATTERN COPYING MATRICES PAPERCUT;
MODEL: F1 BY PATTERN* COPYING MATRICES PAPERCUT;
F1@1;
OUTPUT: SAMPSTAT STANDARDIZED(STDYX) RESIDUAL;## Mplus VERSION 8.4
## MUTHEN & MUTHEN
## 06/10/2021 12:19 PM
##
## INPUT INSTRUCTIONS
##
## TITLE: ONE FACTOR CONFIRMATORY FACTOR ANALYSIS;
## ABSTRACT VISUAL REASONING SCALE - SB4
## DATA: FILE IS "data\AVRS.DAT";
## TYPE IS COVARIANCE;
## NOBSERVATIONS ARE 313;
## VARIABLE: NAMES ARE PATTERN COPYING MATRICES PAPERCUT;
## MODEL: F1 BY PATTERN* COPYING MATRICES PAPERCUT;
## F1@1;
## OUTPUT: SAMPSTAT STANDARDIZED(STDYX) RESIDUAL;
##
##
##
## 1 ERROR(S) FOUND IN THE INPUT INSTRUCTIONS
##
##
##
## ONE FACTOR CONFIRMATORY FACTOR ANALYSIS;
## ABSTRACT VISUAL REASONING SCALE - SB4
##
## SUMMARY OF ANALYSIS
##
## Number of groups 1
## Number of observations 313
##
## Number of dependent variables 4
## Number of independent variables 0
## Number of continuous latent variables 1
##
## Observed dependent variables
##
## Continuous
## PATTERN COPYING MATRICES PAPERCUT
##
## Continuous latent variables
## F1
##
##
## Estimator ML
## Information matrix EXPECTED
## Maximum number of iterations 1000
## Convergence criterion 0.500D-04
## Maximum number of steepest descent iterations 20
##
## Input data file(s)
## data\AVRS.DAT
##
## Input data format FREE
##
##
## SAMPLE STATISTICS
##
##
## SAMPLE STATISTICS
##
##
## Covariances/Correlations/Residual Correlations
## PATTERN COPYING MATRICES PAPERCUT
## ________ ________ ________ ________
## PATTERN 64.000
## COPYING 37.120 64.000
## MATRICES 35.200 32.640 64.000
## PAPERCUT 28.800 32.640 33.280 64.000
##
##
## THE MODEL ESTIMATION TERMINATED NORMALLY
##
##
##
## MODEL FIT INFORMATION
##
## Number of Free Parameters 8
##
## Loglikelihood
##
## H0 Value -4178.740
## H1 Value -4175.037
##
## Information Criteria
##
## Akaike (AIC) 8373.479
## Bayesian (BIC) 8403.449
## Sample-Size Adjusted BIC 8378.075
## (n* = (n + 2) / 24)
##
## Chi-Square Test of Model Fit
##
## Value 7.406
## Degrees of Freedom 2
## P-Value 0.0247
##
## RMSEA (Root Mean Square Error Of Approximation)
##
## Estimate 0.093
## 90 Percent C.I. 0.028 0.169
## Probability RMSEA <= .05 0.117
##
## CFI/TLI
##
## CFI 0.986
## TLI 0.959
##
## Chi-Square Test of Model Fit for the Baseline Model
##
## Value 405.864
## Degrees of Freedom 6
## P-Value 0.0000
##
## SRMR (Standardized Root Mean Square Residual)
##
## Value 0.023
##
##
##
## MODEL RESULTS
##
## Two-Tailed
## Estimate S.E. Est./S.E. P-Value
##
## F1 BY
## PATTERN 5.940 0.430 13.827 0.000
## COPYING 5.978 0.429 13.936 0.000
## MATRICES 5.808 0.432 13.444 0.000
## PAPERCUT 5.319 0.442 12.044 0.000
##
## Variances
## F1 1.000 0.000 999.000 999.000
##
## Residual Variances
## PATTERN 28.513 3.230 8.828 0.000
## COPYING 28.064 3.218 8.721 0.000
## MATRICES 30.065 3.275 9.180 0.000
## PAPERCUT 35.508 3.489 10.177 0.000
##
##
## STANDARDIZED MODEL RESULTS
##
##
## STDYX Standardization
##
## Two-Tailed
## Estimate S.E. Est./S.E. P-Value
##
## F1 BY
## PATTERN 0.744 0.035 21.294 0.000
## COPYING 0.748 0.035 21.586 0.000
## MATRICES 0.727 0.036 20.288 0.000
## PAPERCUT 0.666 0.039 16.860 0.000
##
## Variances
## F1 1.000 0.000 999.000 999.000
##
## Residual Variances
## PATTERN 0.447 0.052 8.604 0.000
## COPYING 0.440 0.052 8.477 0.000
## MATRICES 0.471 0.052 9.042 0.000
## PAPERCUT 0.557 0.053 10.581 0.000
##
##
## R-SQUARE
##
## Observed Two-Tailed
## Variable Estimate S.E. Est./S.E. P-Value
##
## PATTERN 0.553 0.052 10.647 0.000
## COPYING 0.560 0.052 10.793 0.000
## MATRICES 0.529 0.052 10.144 0.000
## PAPERCUT 0.443 0.053 8.430 0.000
##
##
## QUALITY OF NUMERICAL RESULTS
##
## Condition Number for the Information Matrix 0.186E+00
## (ratio of smallest to largest eigenvalue)
##
##
## RESIDUAL OUTPUT
##
##
## ESTIMATED MODEL AND RESIDUALS (OBSERVED - ESTIMATED)
##
##
## Model Estimated Covariances/Correlations/Residual Correlations
## PATTERN COPYING MATRICES PAPERCUT
## ________ ________ ________ ________
## PATTERN 63.796
## COPYING 35.507 63.796
## MATRICES 34.498 34.717 63.796
## PAPERCUT 31.593 31.793 30.890 63.796
##
##
## Residuals for Covariances/Correlations/Residual Correlations
## PATTERN COPYING MATRICES PAPERCUT
## ________ ________ ________ ________
## PATTERN 0.000
## COPYING 1.495 0.000
## MATRICES 0.589 -2.181 0.000
## PAPERCUT -2.885 0.743 2.284 0.000
##
##
## Standardized Residuals (z-scores) for Covariances/Correlations/Residual Corr
## PATTERN COPYING MATRICES PAPERCUT
## ________ ________ ________ ________
## PATTERN 999.000
## COPYING 1.519 999.000
## MATRICES 0.622 -4.255 999.000
## PAPERCUT -3.409 0.634 1.644 999.000
##
##
## Normalized Residuals for Covariances/Correlations/Residual Correlations
## PATTERN COPYING MATRICES PAPERCUT
## ________ ________ ________ ________
## PATTERN 0.000
## COPYING 0.359 0.000
## MATRICES 0.143 -0.539 0.000
## PAPERCUT -0.730 0.183 0.562 0.000
##
##
## Beginning Time: 12:19:13
## Ending Time: 12:19:14
## Elapsed Time: 00:00:01
##
##
##
## MUTHEN & MUTHEN
## 3463 Stoner Ave.
## Los Angeles, CA 90066
##
## Tel: (310) 391-9971
## Fax: (310) 391-8971
## Web: www.StatModel.com
## Support: Support@StatModel.com
##
## Copyright (c) 1998-2019 Muthen & Muthen4.3.2 Calculate coefficient omega \({\omega _u}\)
TITLE: ONE FACTOR CONFIRMATORY FACTOR ANALYSIS;
ABSTRACT VISUAL REASONING SCALE - SB4
CALCULATE COEFFICIENT OMEGA
DATA: FILE IS "data\AVRS.DAT";
TYPE IS COVARIANCE;
NOBSERVATIONS ARE 313;
VARIABLE: NAMES ARE PATTERN COPYING MATRICES PAPERCUT;
MODEL: F1 BY PATTERN* COPYING MATRICES PAPERCUT (la1-la4);
F1@1;
PATTERN COPYING MATRICES PAPERCUT (e1-e4);
MODEL CONSTRAINT:
NEW(omega);
omega=(la1+la2+la3+la4)^2/((la1+la2+la3+la4)^2+e1+e2+e3+e4);
OUTPUT: ## Mplus VERSION 8.4
## MUTHEN & MUTHEN
## 06/10/2021 12:19 PM
##
## INPUT INSTRUCTIONS
##
## TITLE: ONE FACTOR CONFIRMATORY FACTOR ANALYSIS;
## ABSTRACT VISUAL REASONING SCALE - SB4
## CALCULATE COEFFICIENT OMEGA
## DATA: FILE IS "data\AVRS.DAT";
## TYPE IS COVARIANCE;
## NOBSERVATIONS ARE 313;
## VARIABLE: NAMES ARE PATTERN COPYING MATRICES PAPERCUT;
## MODEL: F1 BY PATTERN* COPYING MATRICES PAPERCUT (la1-la4);
## F1@1;
## PATTERN COPYING MATRICES PAPERCUT (e1-e4);
## MODEL CONSTRAINT:
## NEW(omega);
## omega=(la1+la2+la3+la4)^2/((la1+la2+la3+la4)^2+e1+e2+e3+e4);
## OUTPUT:
##
##
##
## 1 ERROR(S) FOUND IN THE INPUT INSTRUCTIONS
##
##
##
## ONE FACTOR CONFIRMATORY FACTOR ANALYSIS;
## ABSTRACT VISUAL REASONING SCALE - SB4
## CALCULATE COEFFICIENT OMEGA
##
## SUMMARY OF ANALYSIS
##
## Number of groups 1
## Number of observations 313
##
## Number of dependent variables 4
## Number of independent variables 0
## Number of continuous latent variables 1
##
## Observed dependent variables
##
## Continuous
## PATTERN COPYING MATRICES PAPERCUT
##
## Continuous latent variables
## F1
##
##
## Estimator ML
## Information matrix EXPECTED
## Maximum number of iterations 1000
## Convergence criterion 0.500D-04
## Maximum number of steepest descent iterations 20
##
## Input data file(s)
## data\AVRS.DAT
##
## Input data format FREE
##
##
##
## THE MODEL ESTIMATION TERMINATED NORMALLY
##
##
##
## MODEL FIT INFORMATION
##
## Number of Free Parameters 8
##
## Loglikelihood
##
## H0 Value -4178.740
## H1 Value -4175.037
##
## Information Criteria
##
## Akaike (AIC) 8373.479
## Bayesian (BIC) 8403.449
## Sample-Size Adjusted BIC 8378.075
## (n* = (n + 2) / 24)
##
## Chi-Square Test of Model Fit
##
## Value 7.406
## Degrees of Freedom 2
## P-Value 0.0247
##
## RMSEA (Root Mean Square Error Of Approximation)
##
## Estimate 0.093
## 90 Percent C.I. 0.028 0.169
## Probability RMSEA <= .05 0.117
##
## CFI/TLI
##
## CFI 0.986
## TLI 0.959
##
## Chi-Square Test of Model Fit for the Baseline Model
##
## Value 405.864
## Degrees of Freedom 6
## P-Value 0.0000
##
## SRMR (Standardized Root Mean Square Residual)
##
## Value 0.023
##
##
##
## MODEL RESULTS
##
## Two-Tailed
## Estimate S.E. Est./S.E. P-Value
##
## F1 BY
## PATTERN 5.940 0.430 13.827 0.000
## COPYING 5.978 0.429 13.936 0.000
## MATRICES 5.808 0.432 13.444 0.000
## PAPERCUT 5.319 0.442 12.044 0.000
##
## Variances
## F1 1.000 0.000 999.000 999.000
##
## Residual Variances
## PATTERN 28.513 3.230 8.828 0.000
## COPYING 28.064 3.218 8.721 0.000
## MATRICES 30.065 3.275 9.180 0.000
## PAPERCUT 35.508 3.489 10.177 0.000
##
## New/Additional Parameters
## OMEGA 0.813 0.017 47.131 0.000
##
##
## QUALITY OF NUMERICAL RESULTS
##
## Condition Number for the Information Matrix 0.508E-01
## (ratio of smallest to largest eigenvalue)
##
##
## Beginning Time: 12:19:14
## Ending Time: 12:19:14
## Elapsed Time: 00:00:00
##
##
##
## MUTHEN & MUTHEN
## 3463 Stoner Ave.
## Los Angeles, CA 90066
##
## Tel: (310) 391-9971
## Fax: (310) 391-8971
## Web: www.StatModel.com
## Support: Support@StatModel.com
##
## Copyright (c) 1998-2019 Muthen & Muthen4.4 Syntax - Mplus - TWo-Factor CFA
4.4.1 An example
TITLE: TWO FACTOR CONFIRMATORY FACTOR ANALYSIS;
ABSTRACT VISUAL REASONING SCALE - SB4
DATA: FILE IS "data\AVRS2F.TXT";
TYPE IS COVARIANCE;
NOBSERVATIONS ARE 313;
VARIABLE: NAMES ARE PATTERN COPYING MATRICES PAPERCUT
QUANT NUMBSER EQUATION;
MODEL: ABSVIS BY PATTERN* COPYING MATRICES PAPERCUT;
QUANTITA BY QUANT* NUMBSER EQUATION;
ABSVIS WITH QUANTITA;
ABSVIS@1; QUANTITA@1;
OUTPUT: SAMPSTAT STANDARDIZED(STDYX) RESIDUAL;## Mplus VERSION 8.4
## MUTHEN & MUTHEN
## 06/10/2021 12:19 PM
##
## INPUT INSTRUCTIONS
##
## TITLE: TWO FACTOR CONFIRMATORY FACTOR ANALYSIS;
## ABSTRACT VISUAL REASONING SCALE - SB4
## DATA: FILE IS "data\AVRS2F.TXT";
## TYPE IS COVARIANCE;
## NOBSERVATIONS ARE 313;
## VARIABLE: NAMES ARE PATTERN COPYING MATRICES PAPERCUT
## QUANT NUMBSER EQUATION;
## MODEL: ABSVIS BY PATTERN* COPYING MATRICES PAPERCUT;
## QUANTITA BY QUANT* NUMBSER EQUATION;
## ABSVIS WITH QUANTITA;
## ABSVIS@1; QUANTITA@1;
## OUTPUT: SAMPSTAT STANDARDIZED(STDYX) RESIDUAL;
##
##
##
## 1 ERROR(S) FOUND IN THE INPUT INSTRUCTIONS
##
##
##
## TWO FACTOR CONFIRMATORY FACTOR ANALYSIS;
## ABSTRACT VISUAL REASONING SCALE - SB4
##
## SUMMARY OF ANALYSIS
##
## Number of groups 1
## Number of observations 313
##
## Number of dependent variables 7
## Number of independent variables 0
## Number of continuous latent variables 2
##
## Observed dependent variables
##
## Continuous
## PATTERN COPYING MATRICES PAPERCUT QUANT NUMBSER
## EQUATION
##
## Continuous latent variables
## ABSVIS QUANTITA
##
##
## Estimator ML
## Information matrix EXPECTED
## Maximum number of iterations 1000
## Convergence criterion 0.500D-04
## Maximum number of steepest descent iterations 20
##
## Input data file(s)
## data\AVRS2F.TXT
##
## Input data format FREE
##
##
## SAMPLE STATISTICS
##
##
## SAMPLE STATISTICS
##
##
## Covariances/Correlations/Residual Correlations
## PATTERN COPYING MATRICES PAPERCUT QUANT
## ________ ________ ________ ________ ________
## PATTERN 64.000
## COPYING 37.120 64.000
## MATRICES 35.200 32.640 64.000
## PAPERCUT 28.800 32.640 33.280 64.000
## QUANT 33.280 31.360 36.480 32.640 64.000
## NUMBSER 34.560 30.080 47.360 39.040 42.240
## EQUATION 21.120 4.480 16.640 17.280 27.520
##
##
## Covariances/Correlations/Residual Correlations
## NUMBSER EQUATION
## ________ ________
## NUMBSER 64.000
## EQUATION 28.160 64.000
##
##
## THE MODEL ESTIMATION TERMINATED NORMALLY
##
##
##
## MODEL FIT INFORMATION
##
## Number of Free Parameters 15
##
## Loglikelihood
##
## H0 Value -7192.765
## H1 Value -7137.604
##
## Information Criteria
##
## Akaike (AIC) 14415.530
## Bayesian (BIC) 14471.723
## Sample-Size Adjusted BIC 14424.148
## (n* = (n + 2) / 24)
##
## Chi-Square Test of Model Fit
##
## Value 110.322
## Degrees of Freedom 13
## P-Value 0.0000
##
## RMSEA (Root Mean Square Error Of Approximation)
##
## Estimate 0.155
## 90 Percent C.I. 0.129 0.182
## Probability RMSEA <= .05 0.000
##
## CFI/TLI
##
## CFI 0.905
## TLI 0.847
##
## Chi-Square Test of Model Fit for the Baseline Model
##
## Value 1047.682
## Degrees of Freedom 21
## P-Value 0.0000
##
## SRMR (Standardized Root Mean Square Residual)
##
## Value 0.060
##
##
##
## MODEL RESULTS
##
## Two-Tailed
## Estimate S.E. Est./S.E. P-Value
##
## ABSVIS BY
## PATTERN 5.532 0.416 13.308 0.000
## COPYING 5.153 0.425 12.134 0.000
## MATRICES 6.514 0.390 16.684 0.000
## PAPERCUT 5.530 0.416 13.301 0.000
##
## QUANTITA BY
## QUANT 6.055 0.401 15.108 0.000
## NUMBSER 7.160 0.376 19.047 0.000
## EQUATION 3.670 0.451 8.146 0.000
##
## ABSVIS WITH
## QUANTITA 0.938 0.024 39.281 0.000
##
## Variances
## ABSVIS 1.000 0.000 999.000 999.000
## QUANTITA 1.000 0.000 999.000 999.000
##
## Residual Variances
## PATTERN 33.191 3.038 10.927 0.000
## COPYING 37.246 3.297 11.296 0.000
## MATRICES 21.357 2.389 8.941 0.000
## PAPERCUT 33.215 3.039 10.930 0.000
## QUANT 27.133 2.638 10.287 0.000
## NUMBSER 12.526 2.232 5.612 0.000
## EQUATION 50.325 4.159 12.101 0.000
##
##
## STANDARDIZED MODEL RESULTS
##
##
## STDYX Standardization
##
## Two-Tailed
## Estimate S.E. Est./S.E. P-Value
##
## ABSVIS BY
## PATTERN 0.693 0.034 20.470 0.000
## COPYING 0.645 0.037 17.333 0.000
## MATRICES 0.816 0.025 32.868 0.000
## PAPERCUT 0.692 0.034 20.450 0.000
##
## QUANTITA BY
## QUANT 0.758 0.029 26.492 0.000
## NUMBSER 0.896 0.021 43.365 0.000
## EQUATION 0.460 0.048 9.556 0.000
##
## ABSVIS WITH
## QUANTITA 0.938 0.024 39.281 0.000
##
## Variances
## ABSVIS 1.000 0.000 999.000 999.000
## QUANTITA 1.000 0.000 999.000 999.000
##
## Residual Variances
## PATTERN 0.520 0.047 11.100 0.000
## COPYING 0.584 0.048 12.158 0.000
## MATRICES 0.335 0.040 8.270 0.000
## PAPERCUT 0.521 0.047 11.106 0.000
## QUANT 0.425 0.043 9.804 0.000
## NUMBSER 0.196 0.037 5.297 0.000
## EQUATION 0.789 0.044 17.851 0.000
##
##
## R-SQUARE
##
## Observed Two-Tailed
## Variable Estimate S.E. Est./S.E. P-Value
##
## PATTERN 0.480 0.047 10.235 0.000
## COPYING 0.416 0.048 8.666 0.000
## MATRICES 0.665 0.040 16.434 0.000
## PAPERCUT 0.479 0.047 10.225 0.000
## QUANT 0.575 0.043 13.246 0.000
## NUMBSER 0.804 0.037 21.682 0.000
## EQUATION 0.211 0.044 4.778 0.000
##
##
## QUALITY OF NUMERICAL RESULTS
##
## Condition Number for the Information Matrix 0.324E-02
## (ratio of smallest to largest eigenvalue)
##
##
## RESIDUAL OUTPUT
##
##
## ESTIMATED MODEL AND RESIDUALS (OBSERVED - ESTIMATED)
##
##
## Model Estimated Covariances/Correlations/Residual Correlations
## PATTERN COPYING MATRICES PAPERCUT QUANT
## ________ ________ ________ ________ ________
## PATTERN 63.795
## COPYING 28.505 63.795
## MATRICES 36.039 33.567 63.795
## PAPERCUT 30.592 28.494 36.024 63.795
## QUANT 31.434 29.278 37.017 31.422 63.794
## NUMBSER 37.173 34.623 43.775 37.159 43.354
## EQUATION 19.054 17.747 22.438 19.046 22.222
##
##
## Model Estimated Covariances/Correlations/Residual Correlations
## NUMBSER EQUATION
## ________ ________
## NUMBSER 63.795
## EQUATION 26.279 63.795
##
##
## Residuals for Covariances/Correlations/Residual Correlations
## PATTERN COPYING MATRICES PAPERCUT QUANT
## ________ ________ ________ ________ ________
## PATTERN 0.000
## COPYING 8.497 0.000
## MATRICES -0.951 -1.031 0.000
## PAPERCUT -1.884 4.042 -2.851 0.000
## QUANT 1.739 1.982 -0.653 1.114 0.001
## NUMBSER -2.724 -4.640 3.434 1.757 -1.249
## EQUATION 1.999 -13.281 -5.851 -1.822 5.210
##
##
## Residuals for Covariances/Correlations/Residual Correlations
## NUMBSER EQUATION
## ________ ________
## NUMBSER 0.001
## EQUATION 1.791 0.000
##
##
## Standardized Residuals (z-scores) for Covariances/Correlations/Residual Corr
## PATTERN COPYING MATRICES PAPERCUT QUANT
## ________ ________ ________ ________ ________
## PATTERN 0.024
## COPYING 3.914 0.020
## MATRICES -1.007 -0.953 0.022
## PAPERCUT -1.294 2.094 -4.118 0.021
## QUANT 1.073 1.129 -0.677 0.704 0.036
## NUMBSER -5.422 -12.552 2.977 1.538 999.000
## EQUATION 0.857 -5.995 -3.691 -0.817 2.534
##
##
## Standardized Residuals (z-scores) for Covariances/Correlations/Residual Corr
## NUMBSER EQUATION
## ________ ________
## NUMBSER 0.035
## EQUATION 1.681 0.019
##
##
## Normalized Residuals for Covariances/Correlations/Residual Correlations
## PATTERN COPYING MATRICES PAPERCUT QUANT
## ________ ________ ________ ________ ________
## PATTERN 0.000
## COPYING 2.038 0.000
## MATRICES -0.231 -0.255 0.000
## PAPERCUT -0.476 0.999 -0.701 0.000
## QUANT 0.428 0.493 -0.157 0.275 0.000
## NUMBSER -0.665 -1.164 0.766 0.416 -0.289
## EQUATION 0.526 -3.674 -1.570 -0.488 1.327
##
##
## Normalized Residuals for Covariances/Correlations/Residual Correlations
## NUMBSER EQUATION
## ________ ________
## NUMBSER 0.000
## EQUATION 0.455 0.000
##
##
## Beginning Time: 12:19:14
## Ending Time: 12:19:14
## Elapsed Time: 00:00:00
##
##
##
## MUTHEN & MUTHEN
## 3463 Stoner Ave.
## Los Angeles, CA 90066
##
## Tel: (310) 391-9971
## Fax: (310) 391-8971
## Web: www.StatModel.com
## Support: Support@StatModel.com
##
## Copyright (c) 1998-2019 Muthen & Muthen4.4.2 Model comparison example - two-factor CFA model across time
TITLE: TWO FACTOR CONFIRMATORY FACTOR ANALYSIS
ACROSS TIME
DATA: FILE IS "data\WERHLE.txt";
TYPE IS COVARIANCE;
NOBSERVATIONS ARE 119;
VARIABLE: NAMES ARE CHANGE1 INFLUEN1 SUGGEST1
CHANGE2 INFLUEN2 SUGGEST2;
MODEL: CONTROL1 BY CHANGE1* INFLUEN1 SUGGEST1;
CONTROL2 BY CHANGE2* INFLUEN2 SUGGEST2;
CONTROL1 WITH CONTROL2;
CONTROL1@1; CONTROL2@1;
OUTPUT: SAMPSTAT STANDARDIZED RESIDUAL;## Reading model: mplus/mplussyntax.out## TWO FACTOR CONFIRMATORY FACTOR ANALYSIS ACROSS TIMEEstimated using ML
## Number of obs: 119, number of (free) parameters: 13
##
## Model: Chi2(df = 8) = 18.648, p = 0.0169
## Baseline model: Chi2(df = 15) = 381.003, p = 0
##
## Fit Indices:
##
## CFI = 0.971, TLI = 0.945, SRMR = 0.038
## RMSEA = 0.106, 90% CI [0.042, 0.169], p < .05 = 0.068
## AIC = 2121.689, BIC = 2157.817TITLE: TWO FACTOR CONFIRMATORY FACTOR ANALYSIS
ACROSS TIME - CORRELATED ERRORS
DATA: FILE IS "data\WERHLE.txt";
TYPE IS COVARIANCE;
NOBSERVATIONS ARE 119;
VARIABLE: NAMES ARE CHANGE1 INFLUEN1 SUGGEST1
CHANGE2 INFLUEN2 SUGGEST2;
MODEL: CONTROL1 BY CHANGE1* INFLUEN1 SUGGEST1;
CONTROL2 BY CHANGE2* INFLUEN2 SUGGEST2;
CONTROL1 WITH CONTROL2;
CONTROL1@1; CONTROL2@1;
CHANGE1 WITH CHANGE2;
INFLUEN1 WITH INFLUEN2;
SUGGEST1 WITH SUGGEST2;
OUTPUT: SAMPSTAT STANDARDIZED RESIDUAL;## Reading model: mplus/mplussyntax.out## TWO FACTOR CONFIRMATORY FACTOR ANALYSIS ACROSS TIME - CORRELATED ERRORSEstimated using ML
## Number of obs: 119, number of (free) parameters: 16
##
## Model: Chi2(df = 5) = 7.75, p = 0.1706
## Baseline model: Chi2(df = 15) = 381.003, p = 0
##
## Fit Indices:
##
## CFI = 0.992, TLI = 0.977, SRMR = 0.029
## RMSEA = 0.068, 90% CI [0, 0.156], p < .05 = 0.31
## AIC = 2116.791, BIC = 2161.257TITLE: TWO FACTOR CONFIRMATORY FACTOR ANALYSIS
ACROSS TIME - PATH CONSTRAINTS
DATA: FILE IS "data\WERHLE.txt";
TYPE IS COVARIANCE;
NOBSERVATIONS ARE 119;
VARIABLE: NAMES ARE CHANGE1 INFLUEN1 SUGGEST1
CHANGE2 INFLUEN2 SUGGEST2;
MODEL: CONTROL1 BY CHANGE1* (1)
INFLUEN1 (2)
SUGGEST1 (3);
CONTROL2 BY CHANGE2* (1)
INFLUEN2 (2)
SUGGEST2 (3);
CONTROL1 WITH CONTROL2;
CONTROL1@1; CONTROL2@1;
OUTPUT: SAMPSTAT STANDARDIZED RESIDUAL;## Reading model: mplus/mplussyntax.out## TWO FACTOR CONFIRMATORY FACTOR ANALYSIS ACROSS TIME - PATH CONSTRAINTSEstimated using ML
## Number of obs: 119, number of (free) parameters: 10
##
## Model: Chi2(df = 11) = 20.206, p = 0.0426
## Baseline model: Chi2(df = 15) = 381.003, p = 0
##
## Fit Indices:
##
## CFI = 0.975, TLI = 0.966, SRMR = 0.055
## RMSEA = 0.084, 90% CI [0.015, 0.141], p < .05 = 0.154
## AIC = 2117.247, BIC = 2145.038TITLE: TWO FACTOR CONFIRMATORY FACTOR ANALYSIS
ACROSS TIME - PATH CONSTRAINTS AND CORRELATED ERRORS
DATA: FILE IS "data\WERHLE.txt";
TYPE IS COVARIANCE;
NOBSERVATIONS ARE 119;
VARIABLE: NAMES ARE CHANGE1 INFLUEN1 SUGGEST1
CHANGE2 INFLUEN2 SUGGEST2;
MODEL: CONTROL1 BY CHANGE1* (1)
INFLUEN1 (2)
SUGGEST1 (3);
CONTROL2 BY CHANGE2* (1)
INFLUEN2 (2)
SUGGEST2 (3);
CONTROL1 WITH CONTROL2;
CONTROL1@1; CONTROL2@1;
CHANGE1 WITH CHANGE2;
INFLUEN1 WITH INFLUEN2;
SUGGEST1 WITH SUGGEST2;
OUTPUT: SAMPSTAT STANDARDIZED RESIDUAL;## Reading model: mplus/mplussyntax.out## TWO FACTOR CONFIRMATORY FACTOR ANALYSIS ACROSS TIME - PATH CONSTRAINTS AND CORRELATED ERRORSEstimated using ML
## Number of obs: 119, number of (free) parameters: 13
##
## Model: Chi2(df = 8) = 9.022, p = 0.3405
## Baseline model: Chi2(df = 15) = 381.003, p = 0
##
## Fit Indices:
##
## CFI = 0.997, TLI = 0.995, SRMR = 0.048
## RMSEA = 0.033, 90% CI [0, 0.116], p < .05 = 0.548
## AIC = 2112.062, BIC = 2148.1914.5 Model Respecification
4.5.1 Lagrange Multiplier test for adding paths
4.5.1.1 R
Modification indices can be requested by adding the argument modindices = TRUE in the summary() call, or by calling the function modindices() directly. The modindices() function returns a data frame which you can sort or filter to extract what you want.
AIRQUALITY <- '
.331
.431 1.160
.406 .847 .898
.216 .272 .312 .268'
cfa.lm.cov <- getCov(AIRQUALITY, names = c("OVERALL", "CLARITY", "COLOR", "ODOR"))
cfa.lm.model <- '
QUALITY =~ OVERALL + CLARITY + COLOR + ODOR'
cfa.lm.fit <- cfa(cfa.lm.model, sample.cov = cfa.lm.cov, sample.nobs = 57, std.lv = TRUE)
summary(cfa.lm.fit, fit.measures = TRUE, standardized = TRUE, rsquare = TRUE, modindices = TRUE)## lavaan 0.6-8 ended normally after 23 iterations
##
## Estimator ML
## Optimization method NLMINB
## Number of model parameters 8
##
## Number of observations 57
##
## Model Test User Model:
##
## Test statistic 16.325
## Degrees of freedom 2
## P-value (Chi-square) 0.000
##
## Model Test Baseline Model:
##
## Test statistic 163.272
## Degrees of freedom 6
## P-value 0.000
##
## User Model versus Baseline Model:
##
## Comparative Fit Index (CFI) 0.909
## Tucker-Lewis Index (TLI) 0.727
##
## Loglikelihood and Information Criteria:
##
## Loglikelihood user model (H0) -180.152
## Loglikelihood unrestricted model (H1) -171.990
##
## Akaike (AIC) 376.304
## Bayesian (BIC) 392.649
## Sample-size adjusted Bayesian (BIC) 367.500
##
## Root Mean Square Error of Approximation:
##
## RMSEA 0.354
## 90 Percent confidence interval - lower 0.209
## 90 Percent confidence interval - upper 0.523
## P-value RMSEA <= 0.05 0.001
##
## Standardized Root Mean Square Residual:
##
## SRMR 0.063
##
## Parameter Estimates:
##
## Standard errors Standard
## Information Expected
## Information saturated (h1) model Structured
##
## Latent Variables:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## QUALITY =~
## OVERALL 0.466 0.063 7.347 0.000 0.466 0.817
## CLARITY 0.921 0.115 7.975 0.000 0.921 0.863
## COLOR 0.882 0.096 9.146 0.000 0.882 0.939
## ODOR 0.351 0.061 5.736 0.000 0.351 0.685
##
## Variances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .OVERALL 0.108 0.025 4.409 0.000 0.108 0.332
## .CLARITY 0.292 0.075 3.885 0.000 0.292 0.256
## .COLOR 0.104 0.049 2.103 0.035 0.104 0.118
## .ODOR 0.140 0.028 4.962 0.000 0.140 0.531
## QUALITY 1.000 1.000 1.000
##
## R-Square:
## Estimate
## OVERALL 0.668
## CLARITY 0.744
## COLOR 0.882
## ODOR 0.469
##
## Modification Indices:
##
## lhs op rhs mi epc sepc.lv sepc.all sepc.nox
## 10 OVERALL ~~ CLARITY 0.196 -0.019 -0.019 -0.108 -0.108
## 11 OVERALL ~~ COLOR 7.630 -0.123 -0.123 -1.159 -1.159
## 12 OVERALL ~~ ODOR 12.542 0.069 0.069 0.558 0.558
## 13 CLARITY ~~ COLOR 12.542 0.341 0.341 1.959 1.959
## 14 CLARITY ~~ ODOR 7.630 -0.097 -0.097 -0.479 -0.479
## 15 COLOR ~~ ODOR 0.196 -0.014 -0.014 -0.115 -0.115Alternatively, use the modindices() function.
Modification indices are printed out for each nonfree (or nonredundant) parameter. The modification indices are supplemented by the expected parameter change (EPC) values (column epc). The last three columns contain the standardized EPC values (sepc.lv: only standardizing the latent variables; sepc.all: standardizing all variables; sepc.nox: standardizing all but exogenous observed variables).
## lhs op rhs mi epc sepc.lv sepc.all sepc.nox
## 10 OVERALL ~~ CLARITY 0.196 -0.019 -0.019 -0.108 -0.108
## 11 OVERALL ~~ COLOR 7.630 -0.123 -0.123 -1.159 -1.159
## 12 OVERALL ~~ ODOR 12.542 0.069 0.069 0.558 0.558
## 13 CLARITY ~~ COLOR 12.542 0.341 0.341 1.959 1.959
## 14 CLARITY ~~ ODOR 7.630 -0.097 -0.097 -0.479 -0.479
## 15 COLOR ~~ ODOR 0.196 -0.014 -0.014 -0.115 -0.115Add a path based on modification indices and theoretical justification. Add one path at a time.
cfa.lm.model.add.covariance <- '
QUALITY =~ OVERALL + CLARITY + COLOR + ODOR
CLARITY ~~ COLOR' #covariance between residuals added
cfa.lm.add.covariance.fit <- cfa(cfa.lm.model.add.covariance, sample.cov = cfa.lm.cov, sample.nobs = 57, std.lv = TRUE)
summary(cfa.lm.add.covariance.fit, fit.measures = TRUE, standardized = TRUE, rsquare = TRUE, modindices = TRUE)## lavaan 0.6-8 ended normally after 22 iterations
##
## Estimator ML
## Optimization method NLMINB
## Number of model parameters 9
##
## Number of observations 57
##
## Model Test User Model:
##
## Test statistic 4.299
## Degrees of freedom 1
## P-value (Chi-square) 0.038
##
## Model Test Baseline Model:
##
## Test statistic 163.272
## Degrees of freedom 6
## P-value 0.000
##
## User Model versus Baseline Model:
##
## Comparative Fit Index (CFI) 0.979
## Tucker-Lewis Index (TLI) 0.874
##
## Loglikelihood and Information Criteria:
##
## Loglikelihood user model (H0) -174.139
## Loglikelihood unrestricted model (H1) -171.990
##
## Akaike (AIC) 366.278
## Bayesian (BIC) 384.666
## Sample-size adjusted Bayesian (BIC) 356.374
##
## Root Mean Square Error of Approximation:
##
## RMSEA 0.241
## 90 Percent confidence interval - lower 0.046
## 90 Percent confidence interval - upper 0.493
## P-value RMSEA <= 0.05 0.052
##
## Standardized Root Mean Square Residual:
##
## SRMR 0.024
##
## Parameter Estimates:
##
## Standard errors Standard
## Information Expected
## Information saturated (h1) model Structured
##
## Latent Variables:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## QUALITY =~
## OVERALL 0.536 0.062 8.640 0.000 0.536 0.940
## CLARITY 0.771 0.129 5.988 0.000 0.771 0.722
## COLOR 0.749 0.109 6.864 0.000 0.749 0.797
## ODOR 0.396 0.060 6.585 0.000 0.396 0.771
##
## Covariances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .CLARITY ~~
## .COLOR 0.255 0.086 2.952 0.003 0.255 0.608
##
## Variances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .OVERALL 0.038 0.029 1.311 0.190 0.038 0.115
## .CLARITY 0.545 0.121 4.490 0.000 0.545 0.478
## .COLOR 0.322 0.081 3.950 0.000 0.322 0.365
## .ODOR 0.107 0.025 4.258 0.000 0.107 0.405
## QUALITY 1.000 1.000 1.000
##
## R-Square:
## Estimate
## OVERALL 0.885
## CLARITY 0.522
## COLOR 0.635
## ODOR 0.595
##
## Modification Indices:
##
## lhs op rhs mi epc sepc.lv sepc.all sepc.nox
## 11 OVERALL ~~ CLARITY 4.141 0.078 0.078 0.548 0.548
## 12 OVERALL ~~ COLOR 4.141 -0.076 -0.076 -0.692 -0.692
## 14 CLARITY ~~ ODOR 4.141 -0.058 -0.058 -0.240 -0.240
## 15 COLOR ~~ ODOR 4.141 0.056 0.056 0.303 0.3034.5.1.2 Mplus
TITLE: AIR QUALITY MODEL
DATA: FILE IS AIRQUALITY.txt;
TYPE IS COVARIANCE;
NOBSERVATIONS ARE 57;
VARIABLE: NAMES ARE OVERALL CLARITY COLOR ODOR;
MODEL: QUALITY BY OVERALL* CLARITY COLOR ODOR;
QUALITY@1;
OUTPUT: STANDARDIZED(STDYX) MODINDICES(3.84);## Mplus VERSION 8.4
## MUTHEN & MUTHEN
## 06/10/2021 12:19 PM
##
## INPUT INSTRUCTIONS
##
## TITLE: AIR QUALITY MODEL
## DATA: FILE IS AIRQUALITY.txt;
## TYPE IS COVARIANCE;
## NOBSERVATIONS ARE 57;
## VARIABLE: NAMES ARE OVERALL CLARITY COLOR ODOR;
## MODEL: QUALITY BY OVERALL* CLARITY COLOR ODOR;
## QUALITY@1;
## OUTPUT: STANDARDIZED(STDYX) MODINDICES(3.84);
##
## *** ERROR in DATA command
## The file specified for the FILE option cannot be found. Check that this
## file exists: AIRQUALITY.txt
##
##
## MUTHEN & MUTHEN
## 3463 Stoner Ave.
## Los Angeles, CA 90066
##
## Tel: (310) 391-9971
## Fax: (310) 391-8971
## Web: www.StatModel.com
## Support: Support@StatModel.com
##
## Copyright (c) 1998-2019 Muthen & MuthenTITLE: AIR QUALITY MODEL WITH ERROR COVARIANCE
DATA: FILE IS AIRQUALITY.txt;
TYPE IS COVARIANCE;
NOBSERVATIONS ARE 57;
VARIABLE: NAMES ARE OVERALL CLARITY COLOR ODOR;
MODEL: QUALITY BY OVERALL* CLARITY COLOR ODOR;
QUALITY@1;
CLARITY WITH COLOR;
OUTPUT: STANDARDIZED(STDYX) MODINDICES(3.84);## Mplus VERSION 8.4
## MUTHEN & MUTHEN
## 06/10/2021 12:19 PM
##
## INPUT INSTRUCTIONS
##
## TITLE: AIR QUALITY MODEL WITH ERROR COVARIANCE
## DATA: FILE IS AIRQUALITY.txt;
## TYPE IS COVARIANCE;
## NOBSERVATIONS ARE 57;
## VARIABLE: NAMES ARE OVERALL CLARITY COLOR ODOR;
## MODEL: QUALITY BY OVERALL* CLARITY COLOR ODOR;
## QUALITY@1;
## CLARITY WITH COLOR;
## OUTPUT: STANDARDIZED(STDYX) MODINDICES(3.84);
##
## *** ERROR in DATA command
## The file specified for the FILE option cannot be found. Check that this
## file exists: AIRQUALITY.txt
##
##
## MUTHEN & MUTHEN
## 3463 Stoner Ave.
## Los Angeles, CA 90066
##
## Tel: (310) 391-9971
## Fax: (310) 391-8971
## Web: www.StatModel.com
## Support: Support@StatModel.com
##
## Copyright (c) 1998-2019 Muthen & Muthen4.5.2 Wald test for dropping paths
4.5.2.1 R
Drop statistical nonsignificant paths one at a time.
POLDEM <- '
6.880
6.253 15.579
5.844 5.840 10.765
6.088 9.504 6.692 11.216
5.065 5.600 4.938 5.706 6.828
5.751 9.386 4.726 7.444 4.980 11.377
5.809 7.535 7.008 7.483 5.822 6.750 10.798
5.670 7.764 5.645 8.012 5.344 8.244 7.594 10.537'
cfa.wald.cov <- getCov(POLDEM, names = paste("Y", 1:8, sep=""))
cfa.wald.model <- '
F1 =~ a*Y1 + b*Y2 + c*Y3 + d*Y4
F2 =~ a*Y5 + b*Y6 + c*Y7 + d*Y8
Y1 ~~ Y5
Y2 ~~ Y6
Y3 ~~ Y7
Y4 ~~ Y8
Y2 ~~ Y4
Y6 ~~ Y8'
cfa.wald.fit <- cfa(cfa.wald.model, sample.cov = cfa.wald.cov, sample.nobs = 102, std.lv = TRUE)
summary(cfa.wald.fit, fit.measures = TRUE, standardized = TRUE, rsquare = TRUE, modindices = TRUE)## lavaan 0.6-8 ended normally after 47 iterations
##
## Estimator ML
## Optimization method NLMINB
## Number of model parameters 23
## Number of equality constraints 4
##
## Number of observations 102
##
## Model Test User Model:
##
## Test statistic 21.121
## Degrees of freedom 17
## P-value (Chi-square) 0.221
##
## Model Test Baseline Model:
##
## Test statistic 627.140
## Degrees of freedom 28
## P-value 0.000
##
## User Model versus Baseline Model:
##
## Comparative Fit Index (CFI) 0.993
## Tucker-Lewis Index (TLI) 0.989
##
## Loglikelihood and Information Criteria:
##
## Loglikelihood user model (H0) -1797.133
## Loglikelihood unrestricted model (H1) -1786.572
##
## Akaike (AIC) 3632.266
## Bayesian (BIC) 3682.140
## Sample-size adjusted Bayesian (BIC) 3622.126
##
## Root Mean Square Error of Approximation:
##
## RMSEA 0.049
## 90 Percent confidence interval - lower 0.000
## 90 Percent confidence interval - upper 0.107
## P-value RMSEA <= 0.05 0.468
##
## Standardized Root Mean Square Residual:
##
## SRMR 0.059
##
## Parameter Estimates:
##
## Standard errors Standard
## Information Expected
## Information saturated (h1) model Structured
##
## Latent Variables:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## F1 =~
## Y1 (a) 2.154 0.195 11.046 0.000 2.154 0.842
## Y2 (b) 2.604 0.275 9.456 0.000 2.604 0.687
## Y3 (c) 2.604 0.243 10.704 0.000 2.604 0.759
## Y4 (d) 2.740 0.240 11.421 0.000 2.740 0.836
## F2 =~
## Y5 (a) 2.154 0.195 11.046 0.000 2.154 0.806
## Y6 (b) 2.604 0.275 9.456 0.000 2.604 0.765
## Y7 (c) 2.604 0.243 10.704 0.000 2.604 0.820
## Y8 (d) 2.740 0.240 11.421 0.000 2.740 0.837
##
## Covariances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .Y1 ~~
## .Y5 0.581 0.314 1.852 0.064 0.581 0.266
## .Y2 ~~
## .Y6 2.072 0.631 3.285 0.001 2.072 0.343
## .Y3 ~~
## .Y7 0.746 0.526 1.418 0.156 0.746 0.184
## .Y4 ~~
## .Y8 0.470 0.390 1.206 0.228 0.470 0.146
## .Y2 ~~
## .Y4 1.408 0.587 2.398 0.017 1.408 0.284
## .Y6 ~~
## .Y8 1.244 0.500 2.487 0.013 1.244 0.316
## F1 ~~
## F2 0.966 0.025 38.699 0.000 0.966 0.966
##
## Variances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .Y1 1.898 0.371 5.116 0.000 1.898 0.290
## .Y2 7.578 1.173 6.463 0.000 7.578 0.528
## .Y3 4.985 0.828 6.017 0.000 4.985 0.424
## .Y4 3.234 0.619 5.227 0.000 3.234 0.301
## .Y5 2.505 0.446 5.616 0.000 2.505 0.351
## .Y6 4.818 0.796 6.056 0.000 4.818 0.415
## .Y7 3.294 0.601 5.483 0.000 3.294 0.327
## .Y8 3.219 0.618 5.206 0.000 3.219 0.300
## F1 1.000 1.000 1.000
## F2 1.000 1.000 1.000
##
## R-Square:
## Estimate
## Y1 0.710
## Y2 0.472
## Y3 0.576
## Y4 0.699
## Y5 0.649
## Y6 0.585
## Y7 0.673
## Y8 0.700
##
## Modification Indices:
##
## lhs op rhs mi epc sepc.lv sepc.all sepc.nox
## 23 F1 ~~ F1 0.106 0.038 1.000 1.000 1.000
## 24 F2 ~~ F2 0.106 -0.038 -1.000 -1.000 -1.000
## 30 F1 =~ Y5 0.666 -0.162 -0.162 -0.061 -0.061
## 31 F1 =~ Y6 0.323 -0.159 -0.159 -0.047 -0.047
## 32 F1 =~ Y7 1.746 0.372 0.372 0.117 0.117
## 33 F1 =~ Y8 0.086 -0.068 -0.068 -0.021 -0.021
## 34 F2 =~ Y1 0.707 0.167 0.167 0.065 0.065
## 35 F2 =~ Y2 0.796 0.250 0.250 0.066 0.066
## 36 F2 =~ Y3 2.773 -0.471 -0.471 -0.137 -0.137
## 37 F2 =~ Y4 0.078 0.065 0.065 0.020 0.020
## 38 Y1 ~~ Y2 0.010 -0.039 -0.039 -0.010 -0.010
## 39 Y1 ~~ Y3 4.064 0.770 0.770 0.250 0.250
## 40 Y1 ~~ Y4 0.812 -0.289 -0.289 -0.117 -0.117
## 41 Y1 ~~ Y6 1.866 0.409 0.409 0.135 0.135
## 42 Y1 ~~ Y7 0.756 -0.290 -0.290 -0.116 -0.116
## 43 Y1 ~~ Y8 0.388 -0.177 -0.177 -0.072 -0.072
## 44 Y2 ~~ Y3 1.110 -0.608 -0.608 -0.099 -0.099
## 45 Y2 ~~ Y5 0.001 -0.014 -0.014 -0.003 -0.003
## 46 Y2 ~~ Y7 0.733 0.418 0.418 0.084 0.084
## 47 Y2 ~~ Y8 0.731 0.509 0.509 0.103 0.103
## 48 Y3 ~~ Y4 0.054 0.106 0.106 0.026 0.026
## 49 Y3 ~~ Y5 0.013 0.046 0.046 0.013 0.013
## 50 Y3 ~~ Y6 2.121 -0.671 -0.671 -0.137 -0.137
## 51 Y3 ~~ Y8 0.876 -0.401 -0.401 -0.100 -0.100
## 52 Y4 ~~ Y5 0.090 0.098 0.098 0.034 0.034
## 53 Y4 ~~ Y6 1.688 0.656 0.656 0.166 0.166
## 54 Y4 ~~ Y7 0.192 -0.171 -0.171 -0.052 -0.052
## 55 Y5 ~~ Y6 0.829 -0.305 -0.305 -0.088 -0.088
## 56 Y5 ~~ Y7 0.466 0.245 0.245 0.085 0.085
## 57 Y5 ~~ Y8 0.364 -0.189 -0.189 -0.067 -0.067
## 58 Y6 ~~ Y7 0.675 -0.338 -0.338 -0.085 -0.085
## 59 Y7 ~~ Y8 2.120 0.567 0.567 0.174 0.1744.5.2.2 Mplus
TITLE: POLITICAL DEMOCRACY MODEL - WALD TEST EXAMPLE
DATA: FILE IS "data\POLDEM.txt";
TYPE IS COVARIANCE;
NOBSERVATIONS ARE 102;
VARIABLE: NAMES ARE Y1-Y8;
MODEL: F1 BY Y1* (1)
Y2 (2)
Y3 (3)
Y4 (4);
F2 BY Y5* (1)
Y6 (2)
Y7 (3)
Y8 (4);
F1@1; F2@1;
F1 WITH F2;
Y1 WITH Y5;
Y2 WITH Y6;
Y3 WITH Y7;
Y4 WITH Y8;
Y2 WITH Y4;
Y6 WITH Y8;
OUTPUT: SAMPSTAT STANDARDIZED(STDYX);## Mplus VERSION 8.4
## MUTHEN & MUTHEN
## 06/10/2021 12:19 PM
##
## INPUT INSTRUCTIONS
##
## TITLE: POLITICAL DEMOCRACY MODEL - WALD TEST EXAMPLE
## DATA: FILE IS "data\POLDEM.txt";
## TYPE IS COVARIANCE;
## NOBSERVATIONS ARE 102;
## VARIABLE: NAMES ARE Y1-Y8;
## MODEL: F1 BY Y1* (1)
## Y2 (2)
## Y3 (3)
## Y4 (4);
## F2 BY Y5* (1)
## Y6 (2)
## Y7 (3)
## Y8 (4);
## F1@1; F2@1;
## F1 WITH F2;
## Y1 WITH Y5;
## Y2 WITH Y6;
## Y3 WITH Y7;
## Y4 WITH Y8;
## Y2 WITH Y4;
## Y6 WITH Y8;
## OUTPUT: SAMPSTAT STANDARDIZED(STDYX);
##
##
##
## 1 ERROR(S) FOUND IN THE INPUT INSTRUCTIONS
##
##
##
## POLITICAL DEMOCRACY MODEL - WALD TEST EXAMPLE
##
## SUMMARY OF ANALYSIS
##
## Number of groups 1
## Number of observations 102
##
## Number of dependent variables 8
## Number of independent variables 0
## Number of continuous latent variables 2
##
## Observed dependent variables
##
## Continuous
## Y1 Y2 Y3 Y4 Y5 Y6
## Y7 Y8
##
## Continuous latent variables
## F1 F2
##
##
## Estimator ML
## Information matrix EXPECTED
## Maximum number of iterations 1000
## Convergence criterion 0.500D-04
## Maximum number of steepest descent iterations 20
##
## Input data file(s)
## data\POLDEM.txt
##
## Input data format FREE
##
##
## SAMPLE STATISTICS
##
##
## SAMPLE STATISTICS
##
##
## Covariances/Correlations/Residual Correlations
## Y1 Y2 Y3 Y4 Y5
## ________ ________ ________ ________ ________
## Y1 6.880
## Y2 6.253 15.579
## Y3 5.844 5.840 10.765
## Y4 6.088 9.504 6.692 11.216
## Y5 5.065 5.600 4.938 5.706 6.828
## Y6 5.751 9.386 4.726 7.444 4.980
## Y7 5.809 7.535 7.008 7.483 5.822
## Y8 5.670 7.764 5.645 8.012 5.344
##
##
## Covariances/Correlations/Residual Correlations
## Y6 Y7 Y8
## ________ ________ ________
## Y6 11.377
## Y7 6.750 10.798
## Y8 8.244 7.594 10.537
##
##
## THE MODEL ESTIMATION TERMINATED NORMALLY
##
##
##
## MODEL FIT INFORMATION
##
## Number of Free Parameters 19
##
## Loglikelihood
##
## H0 Value -1797.133
## H1 Value -1786.572
##
## Information Criteria
##
## Akaike (AIC) 3632.266
## Bayesian (BIC) 3682.140
## Sample-Size Adjusted BIC 3622.126
## (n* = (n + 2) / 24)
##
## Chi-Square Test of Model Fit
##
## Value 21.121
## Degrees of Freedom 17
## P-Value 0.2209
##
## RMSEA (Root Mean Square Error Of Approximation)
##
## Estimate 0.049
## 90 Percent C.I. 0.000 0.107
## Probability RMSEA <= .05 0.468
##
## CFI/TLI
##
## CFI 0.993
## TLI 0.989
##
## Chi-Square Test of Model Fit for the Baseline Model
##
## Value 627.140
## Degrees of Freedom 28
## P-Value 0.0000
##
## SRMR (Standardized Root Mean Square Residual)
##
## Value 0.059
##
##
##
## MODEL RESULTS
##
## Two-Tailed
## Estimate S.E. Est./S.E. P-Value
##
## F1 BY
## Y1 2.154 0.195 11.046 0.000
## Y2 2.604 0.275 9.456 0.000
## Y3 2.604 0.243 10.704 0.000
## Y4 2.740 0.240 11.421 0.000
##
## F2 BY
## Y5 2.154 0.195 11.046 0.000
## Y6 2.604 0.275 9.456 0.000
## Y7 2.604 0.243 10.704 0.000
## Y8 2.740 0.240 11.421 0.000
##
## F1 WITH
## F2 0.966 0.025 38.699 0.000
##
## Y1 WITH
## Y5 0.581 0.314 1.852 0.064
##
## Y2 WITH
## Y6 2.072 0.631 3.285 0.001
## Y4 1.408 0.587 2.398 0.017
##
## Y3 WITH
## Y7 0.746 0.526 1.418 0.156
##
## Y4 WITH
## Y8 0.470 0.390 1.206 0.228
##
## Y6 WITH
## Y8 1.244 0.500 2.487 0.013
##
## Variances
## F1 1.000 0.000 999.000 999.000
## F2 1.000 0.000 999.000 999.000
##
## Residual Variances
## Y1 1.898 0.371 5.116 0.000
## Y2 7.578 1.173 6.463 0.000
## Y3 4.985 0.828 6.017 0.000
## Y4 3.234 0.619 5.227 0.000
## Y5 2.505 0.446 5.616 0.000
## Y6 4.818 0.796 6.056 0.000
## Y7 3.294 0.601 5.483 0.000
## Y8 3.219 0.618 5.206 0.000
##
##
## STANDARDIZED MODEL RESULTS
##
##
## STDYX Standardization
##
## Two-Tailed
## Estimate S.E. Est./S.E. P-Value
##
## F1 BY
## Y1 0.842 0.035 23.959 0.000
## Y2 0.687 0.049 13.953 0.000
## Y3 0.759 0.042 17.962 0.000
## Y4 0.836 0.035 23.989 0.000
##
## F2 BY
## Y5 0.806 0.038 21.139 0.000
## Y6 0.765 0.045 16.911 0.000
## Y7 0.820 0.038 21.860 0.000
## Y8 0.837 0.035 24.001 0.000
##
## F1 WITH
## F2 0.966 0.025 38.699 0.000
##
## Y1 WITH
## Y5 0.266 0.122 2.190 0.029
##
## Y2 WITH
## Y6 0.343 0.085 4.053 0.000
## Y4 0.284 0.098 2.893 0.004
##
## Y3 WITH
## Y7 0.184 0.118 1.553 0.120
##
## Y4 WITH
## Y8 0.146 0.112 1.306 0.192
##
## Y6 WITH
## Y8 0.316 0.099 3.176 0.001
##
## Variances
## F1 1.000 0.000 999.000 999.000
## F2 1.000 0.000 999.000 999.000
##
## Residual Variances
## Y1 0.290 0.059 4.900 0.000
## Y2 0.528 0.068 7.794 0.000
## Y3 0.424 0.064 6.601 0.000
## Y4 0.301 0.058 5.167 0.000
## Y5 0.351 0.061 5.705 0.000
## Y6 0.415 0.069 6.007 0.000
## Y7 0.327 0.062 5.310 0.000
## Y8 0.300 0.058 5.145 0.000
##
##
## R-SQUARE
##
## Observed Two-Tailed
## Variable Estimate S.E. Est./S.E. P-Value
##
## Y1 0.710 0.059 11.979 0.000
## Y2 0.472 0.068 6.976 0.000
## Y3 0.576 0.064 8.981 0.000
## Y4 0.699 0.058 11.994 0.000
## Y5 0.649 0.061 10.569 0.000
## Y6 0.585 0.069 8.456 0.000
## Y7 0.673 0.062 10.930 0.000
## Y8 0.700 0.058 12.000 0.000
##
##
## QUALITY OF NUMERICAL RESULTS
##
## Condition Number for the Information Matrix 0.104E-02
## (ratio of smallest to largest eigenvalue)
##
##
## Beginning Time: 12:19:18
## Ending Time: 12:19:18
## Elapsed Time: 00:00:00
##
##
##
## MUTHEN & MUTHEN
## 3463 Stoner Ave.
## Los Angeles, CA 90066
##
## Tel: (310) 391-9971
## Fax: (310) 391-8971
## Web: www.StatModel.com
## Support: Support@StatModel.com
##
## Copyright (c) 1998-2019 Muthen & Muthen4.5.3 Higher-order factor models
4.5.3.1 R
SC.corr <- '
1.00
0.31 1.00
0.52 0.45 1.00
0.54 0.46 0.70 1.00
0.15 0.33 0.22 0.21 1.00
0.14 0.28 0.21 0.13 0.72 1.00
0.16 0.32 0.35 0.31 0.59 0.56 1.00
0.23 0.29 0.43 0.36 0.55 0.51 0.65 1.00
0.24 0.13 0.24 0.23 0.25 0.24 0.24 0.30 1.00
0.19 0.26 0.22 0.18 0.34 0.37 0.36 0.32 0.38 1.00
0.16 0.24 0.36 0.30 0.33 0.29 0.44 0.51 0.47 0.50 1.00
0.16 0.21 0.35 0.24 0.31 0.33 0.41 0.39 0.47 0.47 0.55 1.00
0.08 0.18 0.09 0.12 0.19 0.24 0.08 0.21 0.21 0.19 0.19 0.20 1.00
0.01 -0.01 0.03 0.02 0.10 0.13 0.03 0.05 0.26 0.17 0.23 0.26 0.33 1.00
0.06 0.19 0.22 0.22 0.23 0.24 0.20 0.26 0.16 0.23 0.38 0.24 0.42 0.40 1.00
0.04 0.17 0.10 0.07 0.26 0.24 0.12 0.26 0.16 0.22 0.32 0.17 0.42 0.42 0.65 1.00'
SC.SDs <- c(1.84, 1.94, 2.07, 1.82, 2.34, 2.61, 2.48, 2.34, 1.71, 1.93, 2.18, 1.94, 1.31, 1.57, 1.77, 1.47)
SC.cov <- getCov(SC.corr, sds = SC.SDs, names = paste("Y", 1:16, sep=""))
HOCFA.1storder.model <-
paste0('F1 =~ ', paste0('Y', 1:4, collapse=' + '), ' \n',
' F2 =~ ', paste0('Y', 5:8, collapse=' + '), ' \n',
' F3 =~ ', paste0('Y', 9:12, collapse=' + '), ' \n',
' F4 =~ ', paste0('Y', 13:16, collapse=' + '))
HOCFA.2ndorder.model <-
paste0('F1 =~ ', paste0('Y', 1:4, collapse=' + '), ' \n',
' F2 =~ ', paste0('Y', 5:8, collapse=' + '), ' \n',
' F3 =~ ', paste0('Y', 9:12, collapse=' + '), ' \n',
' F4 =~ ', paste0('Y', 13:16, collapse=' + '), ' \n',
' F5 =~ NA*F1 + F2 + F3 + F4
F5 ~~ 1*F5')
HOCFA.bifactor.model <-
paste0('F1 =~ ', paste0('Y', 1:4, collapse=' + '), ' \n',
' F2 =~ ', paste0('Y', 5:8, collapse=' + '), ' \n',
' F3 =~ ', paste0('Y', 9:12, collapse=' + '), ' \n',
' F4 =~ ', paste0('Y', 13:16, collapse=' + '), ' \n',
' F5 =~ ', paste0('Y', 1:16, collapse=' + '), ' \n',
' F1 ~~ 0*F2
F1 ~~ 0*F3
F1 ~~ 0*F4
F1 ~~ 0*F5
F2 ~~ 0*F3
F2 ~~ 0*F4
F2 ~~ 0*F5
F3 ~~ 0*F4
F3 ~~ 0*F5
F4 ~~ 0*F5')
HOCFA.1storder.fit <- cfa(HOCFA.1storder.model, sample.cov = SC.cov, sample.nobs = 251, std.lv = TRUE)
HOCFA.2ndorder.fit <- cfa(HOCFA.2ndorder.model, sample.cov = SC.cov, sample.nobs = 251, std.lv = TRUE)
HOCFA.bifactor.fit <- cfa(HOCFA.bifactor.model, sample.cov = SC.cov, sample.nobs = 251, std.lv = TRUE)
summary(HOCFA.1storder.fit, fit.measures = TRUE, standardized = TRUE, rsquare = TRUE) ## lavaan 0.6-8 ended normally after 21 iterations
##
## Estimator ML
## Optimization method NLMINB
## Number of model parameters 38
##
## Number of observations 251
##
## Model Test User Model:
##
## Test statistic 217.311
## Degrees of freedom 98
## P-value (Chi-square) 0.000
##
## Model Test Baseline Model:
##
## Test statistic 1691.802
## Degrees of freedom 120
## P-value 0.000
##
## User Model versus Baseline Model:
##
## Comparative Fit Index (CFI) 0.924
## Tucker-Lewis Index (TLI) 0.907
##
## Loglikelihood and Information Criteria:
##
## Loglikelihood user model (H0) -7583.706
## Loglikelihood unrestricted model (H1) -7475.051
##
## Akaike (AIC) 15243.412
## Bayesian (BIC) 15377.379
## Sample-size adjusted Bayesian (BIC) 15256.915
##
## Root Mean Square Error of Approximation:
##
## RMSEA 0.070
## 90 Percent confidence interval - lower 0.057
## 90 Percent confidence interval - upper 0.082
## P-value RMSEA <= 0.05 0.006
##
## Standardized Root Mean Square Residual:
##
## SRMR 0.058
##
## Parameter Estimates:
##
## Standard errors Standard
## Information Expected
## Information saturated (h1) model Structured
##
## Latent Variables:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## F1 =~
## Y1 1.139 0.112 10.180 0.000 1.139 0.620
## Y2 1.065 0.121 8.794 0.000 1.065 0.550
## Y3 1.744 0.116 15.091 0.000 1.744 0.844
## Y4 1.508 0.102 14.755 0.000 1.508 0.830
## F2 =~
## Y5 1.853 0.130 14.246 0.000 1.853 0.794
## Y6 1.984 0.148 13.447 0.000 1.984 0.762
## Y7 1.936 0.139 13.952 0.000 1.936 0.782
## Y8 1.764 0.133 13.295 0.000 1.764 0.756
## F3 =~
## Y9 1.021 0.107 9.547 0.000 1.021 0.598
## Y10 1.240 0.119 10.447 0.000 1.240 0.644
## Y11 1.713 0.127 13.518 0.000 1.713 0.787
## Y12 1.390 0.116 11.995 0.000 1.390 0.718
## F4 =~
## Y13 0.704 0.084 8.371 0.000 0.704 0.539
## Y14 0.824 0.101 8.140 0.000 0.824 0.526
## Y15 1.424 0.106 13.470 0.000 1.424 0.806
## Y16 1.166 0.088 13.243 0.000 1.166 0.795
##
## Covariances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## F1 ~~
## F2 0.440 0.062 7.110 0.000 0.440 0.440
## F3 0.472 0.063 7.497 0.000 0.472 0.472
## F4 0.217 0.073 2.980 0.003 0.217 0.217
## F2 ~~
## F3 0.640 0.052 12.413 0.000 0.640 0.640
## F4 0.354 0.068 5.214 0.000 0.354 0.354
## F3 ~~
## F4 0.472 0.065 7.263 0.000 0.472 0.472
##
## Variances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .Y1 2.074 0.207 10.003 0.000 2.074 0.615
## .Y2 2.615 0.252 10.381 0.000 2.615 0.698
## .Y3 1.226 0.203 6.031 0.000 1.226 0.287
## .Y4 1.026 0.158 6.490 0.000 1.026 0.311
## .Y5 2.019 0.246 8.206 0.000 2.019 0.370
## .Y6 2.848 0.324 8.777 0.000 2.848 0.420
## .Y7 2.380 0.282 8.432 0.000 2.380 0.388
## .Y8 2.341 0.264 8.871 0.000 2.341 0.429
## .Y9 1.870 0.189 9.878 0.000 1.870 0.642
## .Y10 2.172 0.228 9.510 0.000 2.172 0.586
## .Y11 1.801 0.248 7.266 0.000 1.801 0.380
## .Y12 1.817 0.211 8.622 0.000 1.817 0.485
## .Y13 1.214 0.119 10.179 0.000 1.214 0.710
## .Y14 1.777 0.173 10.247 0.000 1.777 0.724
## .Y15 1.093 0.179 6.106 0.000 1.093 0.350
## .Y16 0.793 0.123 6.428 0.000 0.793 0.369
## F1 1.000 1.000 1.000
## F2 1.000 1.000 1.000
## F3 1.000 1.000 1.000
## F4 1.000 1.000 1.000
##
## R-Square:
## Estimate
## Y1 0.385
## Y2 0.302
## Y3 0.713
## Y4 0.689
## Y5 0.630
## Y6 0.580
## Y7 0.612
## Y8 0.571
## Y9 0.358
## Y10 0.414
## Y11 0.620
## Y12 0.515
## Y13 0.290
## Y14 0.276
## Y15 0.650
## Y16 0.631## lavaan 0.6-8 ended normally after 33 iterations
##
## Estimator ML
## Optimization method NLMINB
## Number of model parameters 36
##
## Number of observations 251
##
## Model Test User Model:
##
## Test statistic 219.475
## Degrees of freedom 100
## P-value (Chi-square) 0.000
##
## Model Test Baseline Model:
##
## Test statistic 1691.802
## Degrees of freedom 120
## P-value 0.000
##
## User Model versus Baseline Model:
##
## Comparative Fit Index (CFI) 0.924
## Tucker-Lewis Index (TLI) 0.909
##
## Loglikelihood and Information Criteria:
##
## Loglikelihood user model (H0) -7584.788
## Loglikelihood unrestricted model (H1) -7475.051
##
## Akaike (AIC) 15241.576
## Bayesian (BIC) 15368.493
## Sample-size adjusted Bayesian (BIC) 15254.368
##
## Root Mean Square Error of Approximation:
##
## RMSEA 0.069
## 90 Percent confidence interval - lower 0.057
## 90 Percent confidence interval - upper 0.081
## P-value RMSEA <= 0.05 0.007
##
## Standardized Root Mean Square Residual:
##
## SRMR 0.060
##
## Parameter Estimates:
##
## Standard errors Standard
## Information Expected
## Information saturated (h1) model Structured
##
## Latent Variables:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## F1 =~
## Y1 0.955 0.100 9.584 0.000 1.139 0.620
## Y2 0.891 0.106 8.390 0.000 1.063 0.549
## Y3 1.461 0.110 13.219 0.000 1.743 0.844
## Y4 1.266 0.097 13.059 0.000 1.510 0.831
## F2 =~
## Y5 1.263 0.132 9.533 0.000 1.862 0.797
## Y6 1.354 0.145 9.306 0.000 1.996 0.766
## Y7 1.307 0.139 9.401 0.000 1.927 0.779
## Y8 1.190 0.130 9.191 0.000 1.755 0.751
## F3 =~
## Y9 0.487 0.117 4.142 0.000 1.023 0.599
## Y10 0.591 0.141 4.205 0.000 1.242 0.645
## Y11 0.812 0.188 4.307 0.000 1.706 0.784
## Y12 0.664 0.155 4.281 0.000 1.395 0.720
## F4 =~
## Y13 0.608 0.075 8.074 0.000 0.704 0.538
## Y14 0.705 0.091 7.792 0.000 0.816 0.521
## Y15 1.238 0.101 12.275 0.000 1.432 0.811
## Y16 1.004 0.083 12.056 0.000 1.162 0.792
## F5 =~
## F1 0.651 0.107 6.086 0.000 0.545 0.545
## F2 1.083 0.181 6.001 0.000 0.735 0.735
## F3 1.848 0.532 3.472 0.001 0.880 0.880
## F4 0.582 0.104 5.611 0.000 0.503 0.503
##
## Variances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## F5 1.000 1.000 1.000
## .Y1 2.075 0.207 10.003 0.000 2.075 0.615
## .Y2 2.619 0.252 10.384 0.000 2.619 0.699
## .Y3 1.231 0.204 6.039 0.000 1.231 0.288
## .Y4 1.019 0.158 6.437 0.000 1.019 0.309
## .Y5 1.987 0.244 8.129 0.000 1.987 0.364
## .Y6 2.802 0.322 8.706 0.000 2.802 0.413
## .Y7 2.411 0.284 8.492 0.000 2.411 0.394
## .Y8 2.374 0.266 8.931 0.000 2.374 0.435
## .Y9 1.867 0.189 9.864 0.000 1.867 0.641
## .Y10 2.168 0.228 9.492 0.000 2.168 0.584
## .Y11 1.824 0.249 7.326 0.000 1.824 0.385
## .Y12 1.804 0.210 8.573 0.000 1.804 0.481
## .Y13 1.214 0.119 10.183 0.000 1.214 0.710
## .Y14 1.789 0.174 10.275 0.000 1.789 0.729
## .Y15 1.070 0.180 5.956 0.000 1.070 0.343
## .Y16 0.801 0.124 6.478 0.000 0.801 0.372
## .F1 1.000 0.703 0.703
## .F2 1.000 0.460 0.460
## .F3 1.000 0.226 0.226
## .F4 1.000 0.747 0.747
##
## R-Square:
## Estimate
## Y1 0.385
## Y2 0.301
## Y3 0.712
## Y4 0.691
## Y5 0.636
## Y6 0.587
## Y7 0.606
## Y8 0.565
## Y9 0.359
## Y10 0.416
## Y11 0.615
## Y12 0.519
## Y13 0.290
## Y14 0.271
## Y15 0.657
## Y16 0.628
## F1 0.297
## F2 0.540
## F3 0.774
## F4 0.253## lavaan 0.6-8 ended normally after 37 iterations
##
## Estimator ML
## Optimization method NLMINB
## Number of model parameters 48
##
## Number of observations 251
##
## Model Test User Model:
##
## Test statistic 157.659
## Degrees of freedom 88
## P-value (Chi-square) 0.000
##
## Model Test Baseline Model:
##
## Test statistic 1691.802
## Degrees of freedom 120
## P-value 0.000
##
## User Model versus Baseline Model:
##
## Comparative Fit Index (CFI) 0.956
## Tucker-Lewis Index (TLI) 0.940
##
## Loglikelihood and Information Criteria:
##
## Loglikelihood user model (H0) -7553.880
## Loglikelihood unrestricted model (H1) -7475.051
##
## Akaike (AIC) 15203.760
## Bayesian (BIC) 15372.982
## Sample-size adjusted Bayesian (BIC) 15220.816
##
## Root Mean Square Error of Approximation:
##
## RMSEA 0.056
## 90 Percent confidence interval - lower 0.042
## 90 Percent confidence interval - upper 0.070
## P-value RMSEA <= 0.05 0.228
##
## Standardized Root Mean Square Residual:
##
## SRMR 0.055
##
## Parameter Estimates:
##
## Standard errors Standard
## Information Expected
## Information saturated (h1) model Structured
##
## Latent Variables:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## F1 =~
## Y1 1.073 0.121 8.898 0.000 1.073 0.584
## Y2 0.713 0.129 5.533 0.000 0.713 0.368
## Y3 1.316 0.121 10.905 0.000 1.316 0.637
## Y4 1.310 0.109 11.982 0.000 1.310 0.721
## F2 =~
## Y5 1.580 0.197 8.016 0.000 1.580 0.677
## Y6 1.709 0.224 7.640 0.000 1.709 0.656
## Y7 0.769 0.173 4.445 0.000 0.769 0.311
## Y8 0.447 0.172 2.593 0.010 0.447 0.191
## F3 =~
## Y9 0.885 0.140 6.322 0.000 0.885 0.519
## Y10 0.838 0.151 5.545 0.000 0.838 0.435
## Y11 0.906 0.155 5.852 0.000 0.906 0.416
## Y12 0.953 0.147 6.468 0.000 0.953 0.492
## F4 =~
## Y13 0.621 0.087 7.119 0.000 0.621 0.475
## Y14 0.802 0.106 7.585 0.000 0.802 0.512
## Y15 1.234 0.109 11.282 0.000 1.234 0.699
## Y16 1.095 0.093 11.783 0.000 1.095 0.746
## F5 =~
## Y1 0.487 0.129 3.777 0.000 0.487 0.265
## Y2 0.795 0.132 6.018 0.000 0.795 0.411
## Y3 1.104 0.136 8.105 0.000 1.104 0.534
## Y4 0.822 0.123 6.695 0.000 0.822 0.453
## Y5 1.265 0.172 7.360 0.000 1.265 0.542
## Y6 1.315 0.194 6.783 0.000 1.315 0.505
## Y7 1.758 0.158 11.147 0.000 1.758 0.710
## Y8 1.825 0.147 12.451 0.000 1.825 0.781
## Y9 0.645 0.120 5.385 0.000 0.645 0.378
## Y10 0.894 0.132 6.757 0.000 0.894 0.464
## Y11 1.437 0.140 10.261 0.000 1.437 0.661
## Y12 1.046 0.130 8.033 0.000 1.046 0.540
## Y13 0.319 0.091 3.489 0.000 0.319 0.244
## Y14 0.218 0.111 1.967 0.049 0.218 0.139
## Y15 0.688 0.120 5.726 0.000 0.688 0.390
## Y16 0.462 0.101 4.554 0.000 0.462 0.315
##
## Covariances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## F1 ~~
## F2 0.000 0.000 0.000
## F3 0.000 0.000 0.000
## F4 0.000 0.000 0.000
## F5 0.000 0.000 0.000
## F2 ~~
## F3 0.000 0.000 0.000
## F4 0.000 0.000 0.000
## F5 0.000 0.000 0.000
## F3 ~~
## F4 0.000 0.000 0.000
## F5 0.000 0.000 0.000
## F4 ~~
## F5 0.000 0.000 0.000
##
## Variances:
## Estimate Std.Err z-value P(>|z|) Std.lv Std.all
## .Y1 1.983 0.212 9.335 0.000 1.983 0.588
## .Y2 2.608 0.248 10.535 0.000 2.608 0.696
## .Y3 1.317 0.198 6.648 0.000 1.317 0.309
## .Y4 0.907 0.175 5.190 0.000 0.907 0.275
## .Y5 1.355 0.428 3.166 0.002 1.355 0.249
## .Y6 2.137 0.523 4.088 0.000 2.137 0.315
## .Y7 2.442 0.274 8.916 0.000 2.442 0.399
## .Y8 1.925 0.269 7.170 0.000 1.925 0.353
## .Y9 1.712 0.218 7.856 0.000 1.712 0.588
## .Y10 2.209 0.243 9.076 0.000 2.209 0.596
## .Y11 1.847 0.235 7.852 0.000 1.847 0.390
## .Y12 1.745 0.228 7.656 0.000 1.745 0.466
## .Y13 1.222 0.120 10.201 0.000 1.222 0.715
## .Y14 1.765 0.176 10.001 0.000 1.765 0.719
## .Y15 1.124 0.181 6.193 0.000 1.124 0.360
## .Y16 0.740 0.135 5.484 0.000 0.740 0.344
## F1 1.000 1.000 1.000
## F2 1.000 1.000 1.000
## F3 1.000 1.000 1.000
## F4 1.000 1.000 1.000
## F5 1.000 1.000 1.000
##
## R-Square:
## Estimate
## Y1 0.412
## Y2 0.304
## Y3 0.691
## Y4 0.725
## Y5 0.751
## Y6 0.685
## Y7 0.601
## Y8 0.647
## Y9 0.412
## Y10 0.404
## Y11 0.610
## Y12 0.534
## Y13 0.285
## Y14 0.281
## Y15 0.640
## Y16 0.656Plot the path diagram
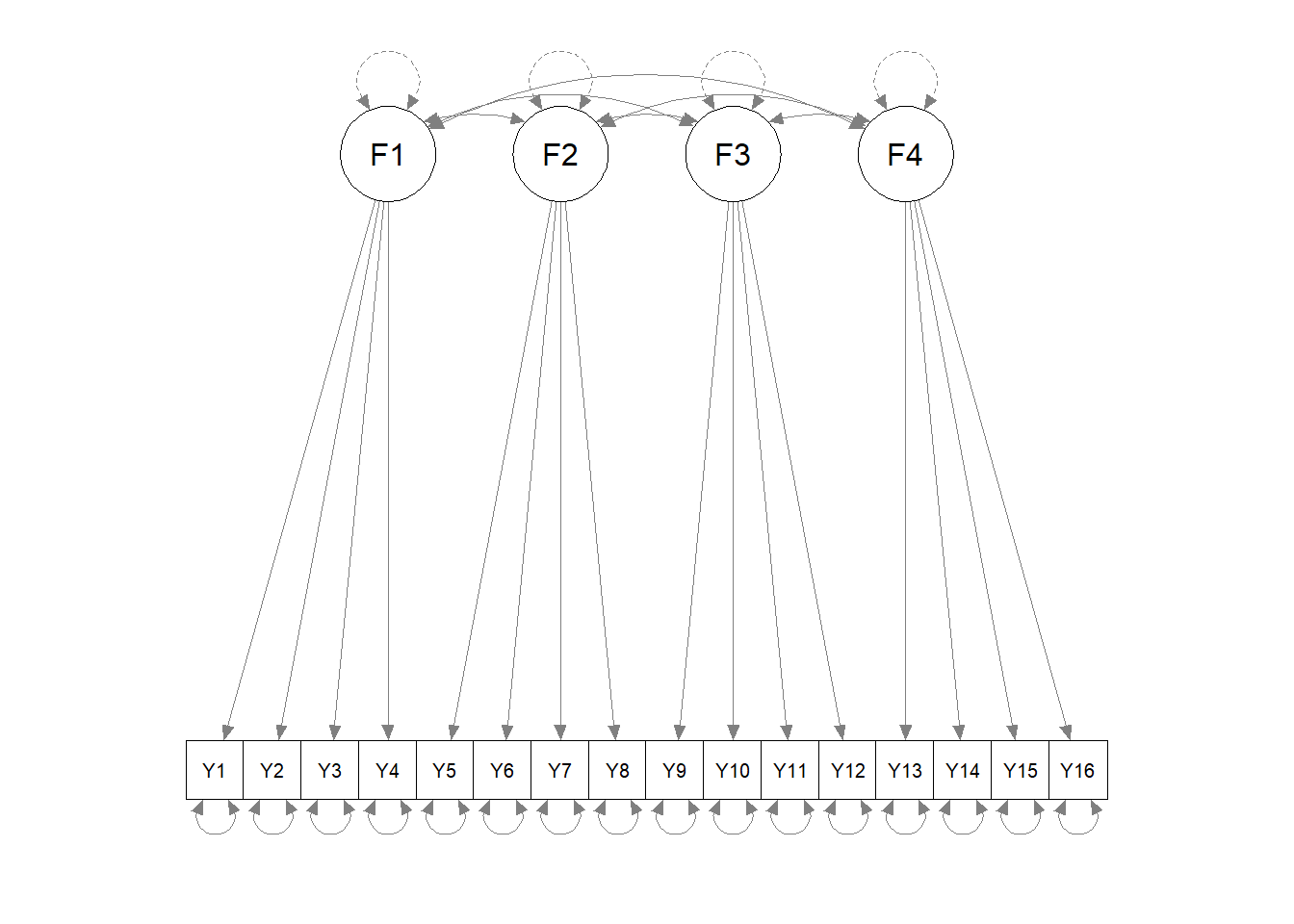
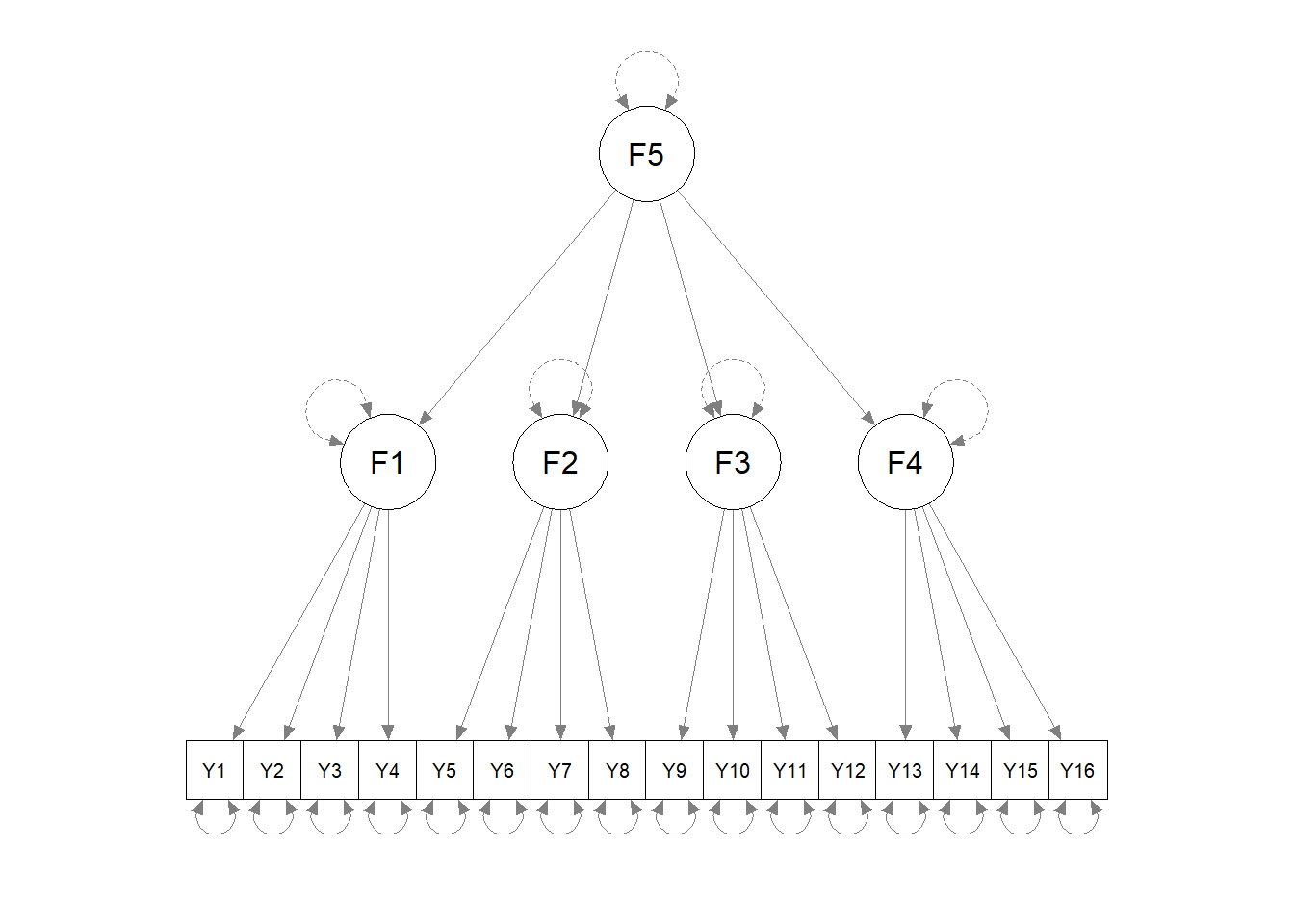
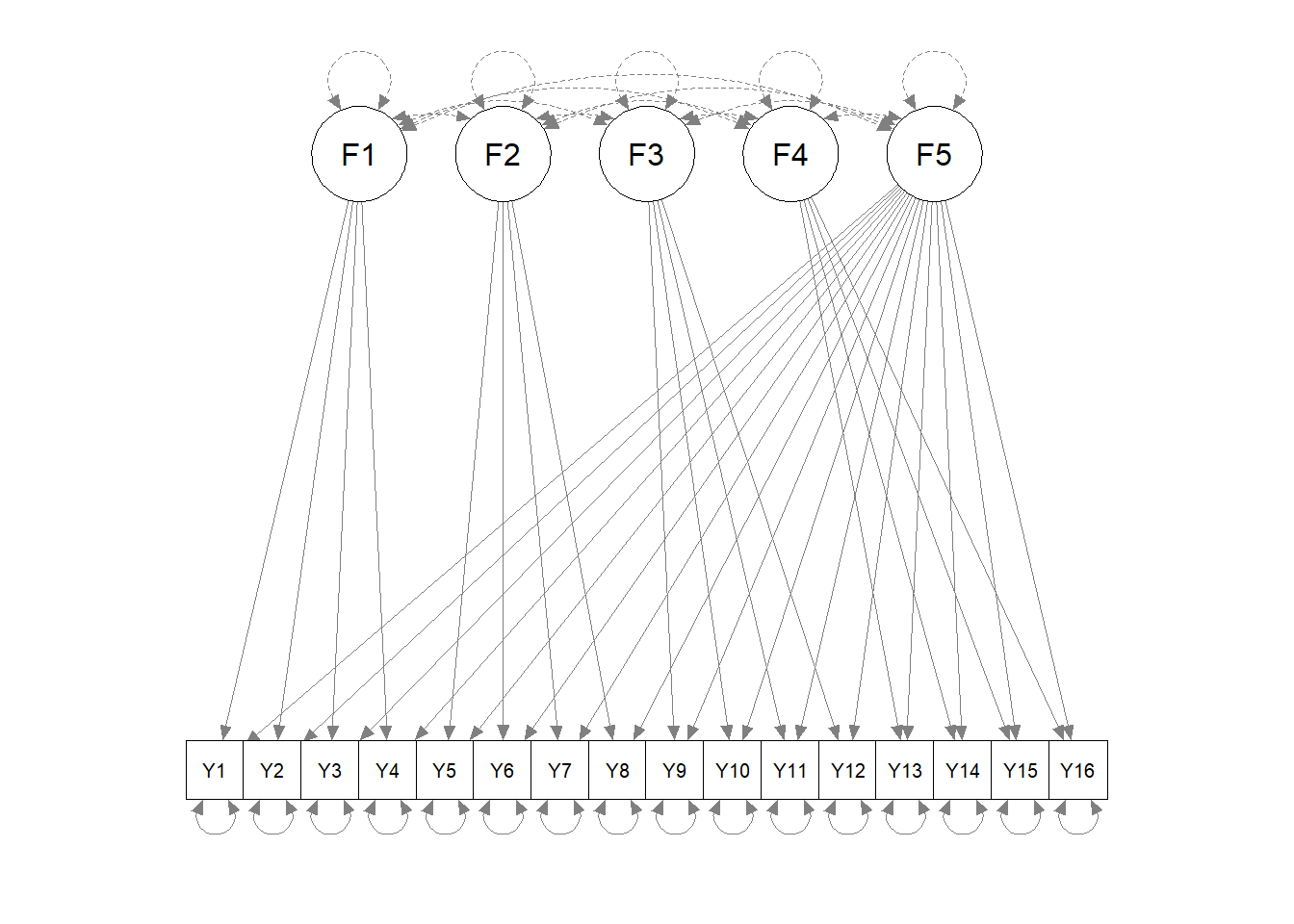
4.5.3.2 Mplus
TITLE: FOUR FACTOR CFA MODEL FOR SELF-CONCEPT;
DATA: FILE IS "data\SC.TXT";
TYPE IS CORRELATION STDEVIATIONS;
NOBSERVATIONS ARE 251;
VARIABLE: NAMES ARE Y1-Y16;
MODEL: F1 BY Y1-Y4;
F2 BY Y5-Y8;
F3 BY Y9-Y12;
F4 BY Y13-Y16;
OUTPUT: STANDARDIZED(STDYX) RESIDUAL;## Reading model: mplus/mplussyntax.out## FOUR FACTOR CFA MODEL FOR SELF-CONCEPT
##
## Estimated using ML
## Number of obs: 251, number of (free) parameters: 38
##
## Model: Chi2(df = 98) = 217.311, p = 0
## Baseline model: Chi2(df = 120) = 1691.802, p = 0
##
## Fit Indices:
##
## CFI = 0.924, TLI = 0.907, SRMR = 0.058
## RMSEA = 0.07, 90% CI [0.057, 0.082], p < .05 = 0.006
## AIC = 15243.412, BIC = 15377.379TITLE: SECOND-ORDER CFA MODEL FOR SELF-CONCEPT;
DATA: FILE IS "data\SC.TXT";
TYPE IS CORRELATION STDEVIATIONS;
NOBSERVATIONS ARE 251;
VARIABLE: NAMES ARE Y1-Y16;
MODEL: F1 BY Y1-Y4;
F2 BY Y5-Y8;
F3 BY Y9-Y12;
F4 BY Y13-Y16;
F5 BY F1* F2-F4;
F5@1;
OUTPUT: STANDARDIZED(STDYX) RESIDUAL;## Reading model: mplus/mplussyntax.out## SECOND-ORDER CFA MODEL FOR SELF-CONCEPT
##
## Estimated using ML
## Number of obs: 251, number of (free) parameters: 36
##
## Model: Chi2(df = 100) = 219.475, p = 0
## Baseline model: Chi2(df = 120) = 1691.802, p = 0
##
## Fit Indices:
##
## CFI = 0.924, TLI = 0.909, SRMR = 0.06
## RMSEA = 0.069, 90% CI [0.057, 0.081], p < .05 = 0.007
## AIC = 15241.576, BIC = 15368.493TITLE: BIFACTOR MODEL FOR SELF-CONCEPT;
DATA: FILE IS "data\SC.TXT";
TYPE IS CORRELATION STDEVIATIONS;
NOBSERVATIONS ARE 251;
VARIABLE: NAMES ARE Y1-Y16;
MODEL: F1 BY Y1-Y4;
F2 BY Y5-Y8;
F3 BY Y9-Y12;
F4 BY Y13-Y16;
F5 BY Y1-Y16;
F1 WITH F2-F5@0;
F2 WITH F3-F5@0;
F3 WITH F4-F5@0;
F4 WITH F5@0;## Reading model: mplus/mplussyntax.out## BIFACTOR MODEL FOR SELF-CONCEPT
##
## Estimated using ML
## Number of obs: 251, number of (free) parameters: 48
##
## Model: Chi2(df = 88) = 157.659, p = 0
## Baseline model: Chi2(df = 120) = 1691.802, p = 0
##
## Fit Indices:
##
## CFI = 0.956, TLI = 0.94, SRMR = 0.055
## RMSEA = 0.056, 90% CI [0.042, 0.07], p < .05 = 0.228
## AIC = 15203.76, BIC = 15372.982